ISSN ONLINE(2319-8753)PRINT(2347-6710)
ISSN ONLINE(2319-8753)PRINT(2347-6710)
Vishal Paramane1, Lalita Admuthe2, Vinayak Sutar3
|
| Related article at Pubmed, Scholar Google |
Visit for more related articles at International Journal of Innovative Research in Science, Engineering and Technology
This study gives a new approach to computer aided detection (CAD) scheme for brain tumor detection and its classification. In the study, large number of MRI images with various types of tumors was selected and examined. The proposed CAD system consist of following steps: 1) MRI image pre-processing; 2) ROI (Region Of Interest) selection; 3) Brain skull edge detection; 4) Tumor Segmentation; 5)Classification of tumor stage. In this study, we have used localised segmentation method for tumor feature extraction and Back Propagation Neural (BPN) Network with Levenberg-Marquardt (LM) algorithm for tumor classification
Keywords |
| Computer aided detection (CAD), Median filter, localised segmentation, Back Propagation Neural (BPN) Network, Levenberg-Marquardt (LM). |
INTRODUCTION |
| Computers were initially used in health care for billing and administrative functions. More recently computers have been used to present clinical information such as laboratory results and pharmacy orders. Many medical informatics researchers believe that the ultimate goal of the “computer-aided diagnosis” should be to advance computerized clinical decision-support. |
| The concept of image processing is an attractive technique for advancement of human perception. Interest in image processing methods stems from two principal application areas: improvement of pictorial information for human interpretation; and processing of image data for storage, transmission, and representation for autonomous machine. It can be applied to analyse images in a large number of domains. The influence and impact of digital images on modern society is tremendous, and image processing is now a critical component in science and technology [7]. The rapid progress in computerized medical image reconstruction, and the associated developments in analysis methods and computer-aided diagnosis, has propelled medical imaging into one of the most important sub-fields in scientific imaging. |
| During the past few decades, with the increasing availability of relatively inexpensive computational resources, computed tomography (CT), magnetic resonance imaging (MRI), Doppler ultrasound, and various imaging techniques based on nuclear emission (PET (positron emission tomography), SPECT (single photon emission computed tomography), etc) have all been valuable additions to the radiologist’s arsenal of imaging tools toward ever more reliable detection and diagnosis of disease. Tomography imaging principles are rooted in physics, mathematics, computer science, and engineering [8]. |
| Medical image processing has experienced dramatic expansion, and has been an interdisciplinary research field attracting expertise from applied mathematics, computer sciences, engineering, statistics, physics, biology and medicine. Computer-aided diagnostic processing has already become an important part of clinical routine. Accompanied by a rush of new development of high technology and use of various imaging modalities, more challenges arise; for example, how to process and analyse a significant volume of images so that high quality information can be produced for disease diagnosis and treatment. |
| To detect an abnormality, the physician usually performs visual scanning of the images of many healthy subjects. The procedure of image scanning lends itself to CAD, since it is a repetitive, burdensome task involving mostly normal images a situation prime for oversight errors [1]. In the case of lesion characterization, complex anatomy, variation in the presentation of benign and malignant states, and the varying abilities of the physicians may result in interpretation errors. CAD systems aim to facilitate lesion detection and characterization tasks by enhancing the physicians’ capabilities and reducing the time required for accurate diagnosis and treatment. |
| Medical image processing is a very broad field; it covers medical signal gathering, image forming, picture processing, and image display to medical diagnosis based on features extracted from images. There are many areas in medical image processing some emphasizing general applicable theory and some focus on specific applications. The objective is to provide a computational output as a second opinion in order to assist physicians in the disease diagnosis and treatment. |
II.RELATED WORK |
| High-resolution MRI determines structural changes in the brain. Manual volumetric methods are currently the gold standard to determine the hippocampal volume, but they are time-consuming. Hippocampal volumetry is the best suited structural biomarker for AD, particularly for early diagnosis and appears to be suitable for risk stratification in mild cognitive impairment cohorts in treatment trials [6]. However, this procedure is still time-consuming and involves a great deal of manual work and therefore is not set to become a routine diagnostic test in the future. The application of Hippocampal volumetry might be further improved in the short-term by implementing semi automated and fully automated analysis procedures [6]. |
| A computer-aided detection (CAD) scheme can be used for pulmonary embolism (PE) detection. For doing this CAD scheme consists of five basic steps: 1) lung segmentation; 2) PE candidate extraction using an intensity mask and tobogganing region growing; 3) PE candidate feature extraction; 4) false-positive (FP) reduction using an artificial neural network (ANN); and 5) a multifeature-based k-nearest neighbour for positive/negative classification [4]. |
| The principles of CAD systems design and development are demonstrated by means of two ways. The first one focuses on the differentiation between symptomatic and asymptomatic carotid atheromatous plaques. For each plaque, a vector of texture and motion features was estimated, which was then reduced to the most robust ones by means of Analysis of Variance (ANOVA). Using fuzzy c-means, the features were then clustered into two classes. Clustering performances of 74%, 79%, and 84% were achieved for texture only, motion only, and combinations of texture and motion features, respectively [1]. |
| The second CAD system supports the diagnosis of focal liver lesions and is able to characterize liver tissue from Computed Tomography (CT) images as normal, hepatic cyst, hemangioma, and hepatocellular carcinoma. Five texture feature sets were extracted for each lesion, while a genetic algorithm based feature selection method was applied to identify the most robust features. The selected feature set was fed into an ensemble of neural network classifiers [1]. |
| The advantage of hierarchical self organizing map and fuzzy c means algorithms are used to classify the image layer by layer. The lowest level weight vector is achieved by the abstraction level. We have also achieved a higher value of tumor pixels by this neuro-fuzzy approach. The multilayer segmentation results of the neuro fuzzy are shown to have interesting consequences from the viewpoint of clinical diagnosis [2]. |
| There are several techniques for image recognition. Among those methods, application of soft computing models on digital image has been considered to be an approach for a better result. The main objective of the present work is to provide a new approach for image recognition using Artificial Neural Networks [3]. Soft Computing is an emerging field built up of latest techniques like fuzzy logic, artificial neural networks, evolutionary computation and machine learning. Each soft computing technique can be applied to produce solutions to any problem that is too complex or noisy to tackle with conventional methods. Artificial Neural Network has been one of the recent development tools that are inspired from biological neural networks. The main advantage of this new powerful tool is to use its capacity to solve problems that are not very easy to be solved by traditional computing methods [3]. |
| The Region of Interest both normal & abnormal appearing in medical image may be defined using manual or semiautomatic methodologies. These methodologies are used according to the application requirement [1]. In brain tumor detection system the part of tumor in scanned brain image is region of interest [9] [12]. |
| Automated extraction of pulmonary anatomy provides a foundation for computerized analysis of computed tomography (CT) scans of the chest. A completely automatic method is presented to segment the lungs, lobes and pulmonary segments from volumetric CT chest scans. The method starts with lung segmentation based on region growing and standard image processing techniques. Next, the pulmonary fissures are extracted by a supervised filter. Subsequently the lung lobes are obtained by voxel classification where the position of voxels in the lung and relative to the fissures is used as features. Finally, each lobe is subdivided in its pulmonary segments by applying another voxels classification that employs features based on the detected fissures and the relative position of voxels in the lobe [10]. |
| Segmentation is an important process to extract information from complex medical images. Image Segmentations refers to the process of partitioning an image into groups of pixels which are homogenous with some criteria. Segmentation algorithms are area oriented instead of pixel oriented. The result of segmentation is the splitting up of the image into connected areas. Thus segmentation is concerned with dividing an image in to meaningful regions [11]. The one of the most common problem of pattern recognition in medical image analysis is the classification of a set into proper class. The extracted features classification can be supervised or unsupervised. In the supervised classification, the feature set is identified as a member of a predefined class, while in unsupervised classification, the feature set pattern is assigned to an unknown class [11]. |
III.SYSTEM BLOCK DIAGRAM |
| The system consists of following steps |
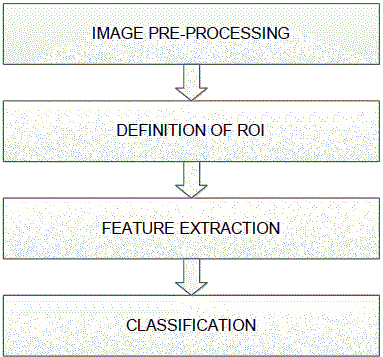 |
ï· Image pre-processing: |
| By image pre-processing we improved the quality of data through the application of methods for denoising. Median filter is used for this. |
Median filter |
| In median filtering, the neighbouring pixels are ranked according to brightness (intensity) and the median value becomes the new value for the central pixel. |
| Median filters can do an excellent job of rejecting certain types of noise, in particular, “shot” or impulse noise in which some individual pixels have extreme values. In the median filtering operation, the pixel values in the neighbourhood window are ranked according to intensity, and the middle value (the median) becomes the output value for the pixel under evaluation. |
| The best known order-statistics filter is the median filter, which replaces the value of a pixel by the median of the gray levels in the neighbourhood of that pixel- |
| The original value of the pixel is included in the computation of the median. |
ï· Definition of ROI and feature extraction: |
| The MRI image of brain consists of different features, to detect the tumor we have to collect similar feature attributes together. For this the particular space in brain image is to be selected which is nothing but the ROI (Region of Interest).In this study we have used computer mouse to specify ROI. This selected area is used to generate binary mask of the region. |
| After defining ROI the binary mask generated is used for feature extraction. Segmentation of image is required for feature extraction. Image segmentation is done by localized segmentation method. |
Localized Segmentation Method |
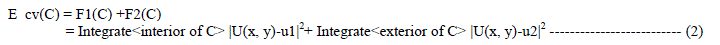 |
ï· Classification: |
| Along with tumor detection classification of tumor stage is essential for effective diagnosis system. For this various methods are available but in this study we are using artificial neural network approach. |
| For classification of tumor stage the back propagation neural network is used. Training a network by back-propagation involves three stages: |
| 1) The feed-forward of the input training pattern |
| 2) The back-propagation of the associated error |
| 3) The adjustment of the weights. |
| The ANN comprised three layers (one input layer, one hidden layer, and one output layer) trained by back propagation. In proposed method back propagation feed forward neural network with Levenberg-Marquardt algorithm is used. |
Levenberg-Marquardt Algorithm (LM) |
| For LM algorithm, the performance index to be optimized is defined as |
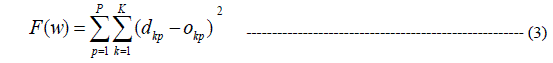 |
| Where e is the error vector comprising the error for all the training examples. When training with the LM method, the increment of weights Δw can be obtained as: |
| Where J is the Jacobian matrix, μ is the learning rate which is to be updated using the β depending on the outcome. In Particular, μ is multiplied by decay rate β (0<β<1) whenever F (w) decreases, whereas μ is divided by β whenever F (w) increases in a new step. |
IV. RESULTS |
| For this method we tested various MRI brain images, the results of pre-processing, ROI, Segmentation is given in this paper. Before segmentation we have measured skull area of each MRI image which is given in Table I. Application of segmentation gave us tumor pixels & tumor area. By using tumor area & skull area we obtained proportional area .All the areas and tumor stages are represented in Table 1. |
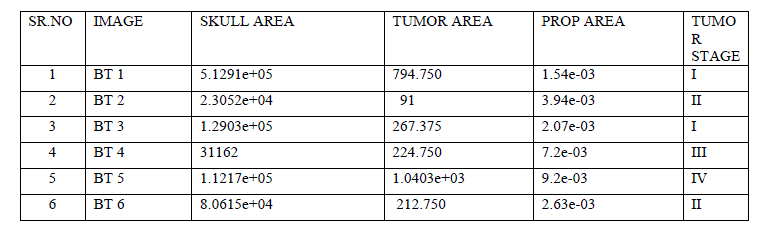 |
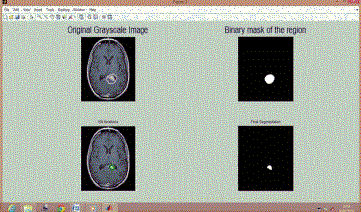
| A number of original and test images have been taken and processed. The results for selected brain MRI images are shown in Fig 1 to Fig 6. |
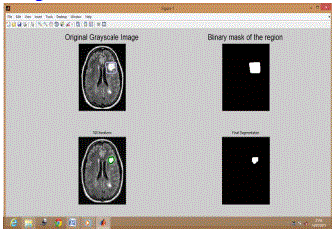
| Figure 1 shows original gray scale tumored brain image (BT 1) in 1st window and selected binary mask of ROI (region of interest) in 2nd window. The processing of segmentation is shown in 3rd window while final segmented tumor is shown in 4th window. |
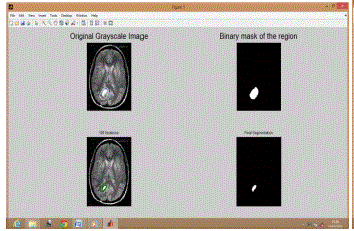
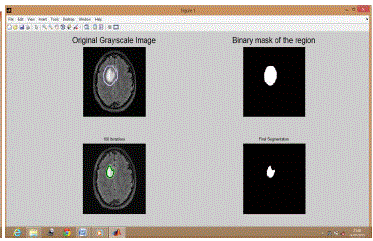
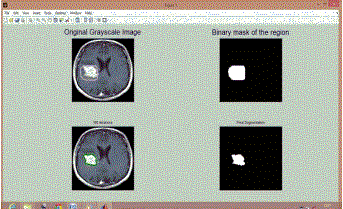
| Similarly Figure 2 shows results for BT 2, Figure 3 shows results for BT 3, Figure 4 shows results for BT 4, Figure 5 shows results for BT 5 and Figure 6 shows results for BT 6. |
| For the process, the binary mask gets applied on original image for given no. of iterations. Here in this method we have taken 100 iterations. The results obtained are improved for each iteration. After 100 iterations we got most accurate results. |
 |
| The proportional area for large no. of MRI brain images is calculated & used to train the neural network. After training, the network is used for testing and validation. |
| Figure 7 shows the process of training the neural network. To train the network we have selected two hidden layers and 29 iterations. The execution time for training was 0:00:02 sec. |
 |
| After training, the network was used for testing. Once the network is trained we can use the network for any value of proportional area. For this, we have to give proportional area in command window. For the given proportional area the network gives certain weight values which are used to find stage of tumor. Figure 8 shows the command window to give proportional area and the same window displays tumor stage. |
 |
| Figure 9 shows the training performance of neural network where x-axis is used for epoch & y-axis for MSE(mean square error).Blue, Red & Green colours shows training, testing & validation performance respectively. At 23rd epoch we get best validation performance. |
 |
| Figure 11 represent Regression of network, 1st window shows training regression, 2nd window shows validation regression & 3rd window shows testing regression. The regression of all is shown in 4th window. |
 |
| Fig. 11 Neural Network Training Regression |
V. CONCLUSION |
| We developed and tested a new CAD scheme for brain tumor detection & classification of tumor stage. To improve the system we tested & examined different MRI brain images. The study showed that the tumor detection & classification can be done effectively with the proposed method. By taking MRI images of a patient at regular interval we can also predict the growth rate of tumor which is very difficult to find without CAD system. The presented CAD system gives 2nd opinion to doctors for accurate detection of tumor & its stages. |
References |
|