ISSN ONLINE(2319-8753)PRINT(2347-6710)
ISSN ONLINE(2319-8753)PRINT(2347-6710)
| Dr.A.Kannan Professor, Department of MCA, K.L.N. College of Engineering, Madurai, India |
| Related article at Pubmed, Scholar Google |
Visit for more related articles at International Journal of Innovative Research in Science, Engineering and Technology
The Content Based Image Retrieval (CBIR) is a popular and powerful technology which is designed to retrieve the desired target image from the large collection of images based on the contents of the given query image. In this paper, the CBIR is designed to support the medical field toretrieve similar and dissimilar conditioned Magnetic Resonance (MR) Images to aid for analysing the condition of brain tumour of a patient using Hybrid K-Nearest NeighbourSupport Vector Machine (HKNNSVM) and Nearest Neighbour (NN) algorithms. All we aware that the field of medical plays a vital role in every country and day to day improvements are being concentrated at regular intervals to save the life of the patients from very crucial diseases. Especially, Magnetic Resonance Image (MRI)dominates medical field to certain extend to identify and examine the patient’s critical problems in an effective and efficient manner. Every day, large numbers of MR Images are being generated and stored in the image database. If these images are processed in a right way, it will reveal useful information to the physicians to take immediate suitable remedial actions for the patient concerned very earlier.
Keywords |
| HKNNSVM, NN, GLCM, Entropy, Cluster Prominence |
1.0 Problem Description |
| The field of multimedia and image processing are nowadays surviving in the age of very advancement technologies today. The Content Based Image Retrieval (CBIR) plays an important part of image processing technique and becomes an emerging technology in the research areas. Processing potential digital image data is a major activity in all problems today. Many users are seeking suitable applications to process and retrieve the image data for the beneficial of their business concerned. They want to locate their desired images in large and varied collections. But, it is highly complicated to locate such images in a huge collection of set. So, image retrieval is the fastest developing and challenging research area with regard to both still and moving images. |
| Thus, the CBIR systems pay attention to image contents called ‘features’. The CBIR extracts features of images to index images with little interventions of human. The feature extraction techniques are classified in various formats such as color, texture and shape. However, the texture retrieval is the most powerful and optimal technique recommended by every researcher in the field of image processing in this present scenario. The CBIR seems to be highly beneficial technique not only for the management of large numbers of image data, but also supports to various fields such as research, clinical medicine, education and visual technology. CBIR is aimed to retrieve desired images based on the similarity measurements. The characteristics of these similarity measurements include intensity, color, texture, size, location and so on. |
| Many CBIR system prototypes have been proposed and few are used as commercial systems (John Eakins et al, 1999). Though, a few of them became commercial products, most of the CBIR systems were designed as research prototype and being developed in universities and research laboratories (Ricardo da Silva Torres et al, 2006). It is aimed to search image databases for specific images that are similar to a given query image. It is necessary to develop new techniques to support effective searching and browsing of large digital libraries based on automatically derived imagery features (Ordonez, C. et al, 1999). Especially, CBIR is focused enormously in medical field in this present scenario. |
| There are two popular diagnostic imaging techniques Computed Tomography (CT) scan and Magnetic Resonance Imaging (MRI) are being implemented in the field of medical to diagnose any abnormal changes in tissues and organs for the early recover of the patients. Among these, MRI is very useful technique to locate the size of the brain tumor very efficiently. MRI is a kind of test in which a magnetic field and the radio wave energy pulses are combined to make pictures of organs and structures inside the body.This creates good contrast between the various soft tissues of the body by which the physicians can move to the right locations to identify the problematic tissues especially in brain, heart, cancer and muscles(Ahmed KHARRAT et al, 2010).The Hybridized KNNSVM algorithm is being effectively used to identify the condition of the brain tumor in human brain. It is highly feasible to process MR Images to classify the type of brain tumor of a person based on the GLCM texture as described in (Kannan, A et al, 2013) |
| K-Nearest Neighbour (K-NN) is one of the supervised learning methods that can be used in many applications in the field of data mining and statistical classification and pattern recognition. This is based on learning by correlation. That is, the given test tuple is to be compared with training tuples that are similar to it. The training tuples that already have been described with ‘n’ number of attributes. The tuples are represented as individual points in an n-dimensional space. When an unknown tuple is received, the K-NN classifier tries to search the pattern space for the ‘k’ training tuples that are closest to the given unknown tuple. The ‘K-nearest neighbours’ are called as ‘k’ training tuples for the given unknown tuple (Jiawei Han et al, 2012). |
| SVM acts as a binary linear classification model which takes input data and predicts for each input data to identify which of two supervised classes the input is a member of which. In other word, SVM classification algorithm builds a model which tries to predict whether the type of given input data belongs to one of the two training categories or not (Jiawei Han et al, 2012)? This is based on the principles of Structural Risk Minimization (SRM) from statistical learning theory (Christopher J.C. Burges, 1998). SRM is an inductive principle for model selection used from learning finite data set and provides a method for controlling the generalization ability learning machines that uses a small size training data (Lam Hong, Lee et al, 2010). The implementation of SRM helps to seek an optimal hyper plane and which guarantees the lowest classification error. |
| It is known that several MR images are being generated in routine processes in the field of medicine for brain tumour. Physicians may want to compare the existing concluded identified diseases with newly arrived patient’s MR Image, so that they can decide suitable remedial actions early for the same. The HKNNSVMdesigned in the work (Kannan, A et al, 2013) is very useful to identify the type of the brain tumour from a patient’s MR Image. However, physicians may wish to classify the existing identified MR images to retrieve the details of similar conditioned MR images such as collections of ‘benign’ conditioned images, ‘Malignant’ conditioned images otherwise heterogeneous collections. In connection with this, a CBIR system is designed in association with the concepts of image mining to fulfil the above said needs. Here, the Nearest- Neighbour (NN) method is implemented in addition to the proposed hybridized KNN SVM method discussed in (Kannan, A et al, 2013) . |
| Hence, the CBIR system is aimed to retrieve either homogenous or heterogeneous tumour conditioned MR images from the database based on the user’s need. The NN method is used to retrieve heterogeneous MR Images from the database, whereas the HKNNSVM algorithm is used to retrieve homogenous MR images from the database. Consequently, the medical staff can compare the results of the given query MR image with the pre-existing tumour conditioned images and necessary remedial solutions can easily be identified to save the patients very earlier. |
2.0 Proposed Solution |
| In this paper, there are two kinds of levels such as training and testing process. The MR images have been collected in the training set in the form of gray scale format. The images are collected from DICOM. The DICOM has already concluded the results of the categories of MR images concerned based on certain criteria. The Grey Level Co-occurrence Matrix (GLCM) has to be calculated from those images to identify texture contents later. The GLCM is used to extract second order statistics from an image. GLCMs have been used very successfully for texture calculations (Yixin Chen et al, 2005) and the GLCM is also used to identify the images in rotation invariants. Since images are collected in different dimension variants, these calculations will be very useful to classify the images in a right way. The GLCM will provide the information about the positions of pixels those have similar gray level values. The matrix will be in the form of two-dimensional array ‘C’ in which the possible image values will be defined as rows and columns. Then, the texture features will be extracted from the collected stored supervised MR images (training) based on the values of the co-occurrence matrix and those features will be kept in a database for future processes. |
| The patient’s query MR image will be received from the user. The GLCM will be calculated from the given query image. A number of texture features may be extracted from the GLCM (Robert M. Haralick et al, 1973). As mentioned earlier, the 12 prominent features such as ‘Contrast’, ‘Correlation’, ‘Cluster Prominence’, ‘Cluster Shade’, ‘Dissimilarity’, ‘Energy’, ‘Entropy’, |
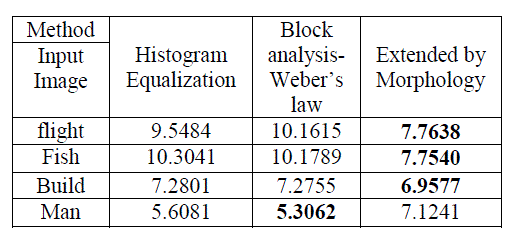
| ‘Homogeneity’, ‘Homogeneity Probability’, ‘Maximum Probability’, ‘Sum of Squares’ and ‘Auto Correlation’ are to be extracted from the query image. The sample features extracted from the images are shown in the Table 2.0. |
 |
| Further, the dimensionality of the data can be reduced by selecting only the subset of measured features to this research work in order to speed up the execution. This is called as Feature Selection. The selection criteria will be usually involved in minimizing specific measure of predictive error for models fit to different subsets. Suitable algorithms are designed to search for a subset of predictors that will optimize the model’s measured responses, subject to constraints such as required or excluded features and the size of the subset (Mathworks R 2012). Thus, the SFS selects the subset of features that can support the best performance on considering the parameters accuracy and computation time. |
| Here, the features “Cluster Prominence” and “Entropy” are received as optimized features by applying SFS. Then, the options can be selected by the users to process their query image either by NN method or Hybrid method. If NN method is selected, the heterogeneous MR Images will be retrieved from the database to the view of human users. If hybrid is selected, the processes which are depicted in the block diagram will be followed to retrieve homogenous MR images as explained in (Kannan, A et al, 2013). The Figure 2.0 shows the block diagram of the entire proposed CBIR system for MRI diagnosis. |
2.1 Implementation and Result Details |
| The query MR image will be selected by the user from the database. Then, the user can either select Nearest Neighbour (NN) method or the hybridized KNNSVM method discussed in the work (Kannan, A et al, 2013) willretrieve nearest images from the database. The user can specify the number of required nearest images such as 5, 10, 15 and 20. Once the number is selected, the neighbours will be selected and the neighbour IDs of each sample will be displayed to the user for reference. In this research work, 150 MR Images have been taken for testing purpose. |
 |
| As pointed out earlier, the NN method will retrieve heterogeneous tumour categorized images from the database. The NN method acts as similar as the KNN method. The NN method in other terms called as proximity search and closest search. This is an optimization problem used to find closet neighbours in the given metric space. A feature space will be formed based on the prominent features of the given images. The query image will be processed to obtain the concerned prominent features and it will be located in the feature space. The distance between given query image and the training set samples will be computed using Euclidean Distance method and the result will be sorted out. The top concerned numbers of images will be picked out as the result. |
| Here, image25 has been given as a query image for the NN classification. The figures 2.1 and 2.2 show the neighbours locations of the given query image and its corresponding distance measurement graph of its neighbour images in the NN classification. It just locates the different neighbours of the given query image. The rounded circle with ‘X’ indicates the given query image. The distance measurements of each neighbour are sorted out and displayed in the distance measurement graph. The distance measurement graph is a bar graph used to identify the distanced level of each neighbour. Here, the choice for number of neighbours selected is 20. The distance measurement graph shows the distances of each MR image. Finally, the CBIR system retrieves discrete categories of tumour formation of MR images which are clearly shown in Figure 2.3. Hence, the physician can compare these results with the given query image in order to take certain remedial actions for the identified problem. |
 |
| Next, the hybrid KNNSVMM has been employed for the given query image25 and it classifies the given query image. The procedures for processing hybrid are similar to the NN method as discussed above. The results of the hybrid algorithm are presented from the Figures 2.4 to 2.6. This hybrid method will retrieve either of ‘Benign’, ‘Normal’ and ‘Malignant’ formation of images from the database according to the given category ofquery image. The given query MR image25 belongs to ‘Benign’ category. So, the hybrid algorithm here has retrieved all the similar conditioned ‘Benign’ categorized MR images from the database according to the classification procedure of the hybrid algorithm discussed in the paper (Kannan, A et al, 2013). Secondly, the query image25 is again applied. The Green pixels show the category of ‘Benign’ which are very close to the given query image25. |
 |
3.0 State of the Art |
| (Aditi P. Killedar et al, 2012) have developed a hybrid technique for medical decision support system to detect tumor in human brain using GLCM, SVM and Principle Component Analysis (PCA). This system was developed to analysis images with tumor and images of multiple sclerosis. |
| There are few more hybrid techniques developed for detection of brain tumor based on texture features extraction, reduction and classification. The systems are developed using the hybrid techniques such as i) Discrete Wavelet Transform(DWT)+ Principal Components Analysis (PCA)+ Artificial Neural Networks(ANN), ii) Discrete Wavelet Transform (DWT)+ Principal Components Analysis(PCA)+ KNearest Neighbors(K-NN), iii) Discrete Wavelet Transform(DWT)+ Self Organization Map (SOM). |
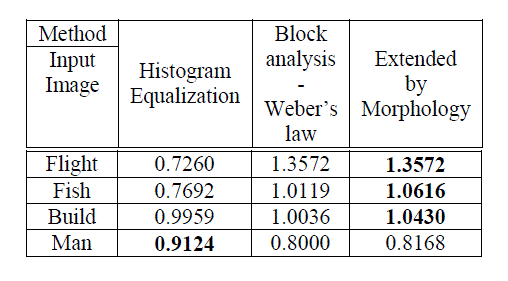
| The proposed CBIR system using hybrid KNNSVM+NN produces high classification accuracy and low computation. This is due to feature reduction and the hybrid concept discussed earlier in(Kannan, A et al, 2013).The Table 3.0 and the Figure 3.0 depict the time consumed by each categorized query images such as ‘Normal’, ‘Benign’, and ‘Malignant’ to retrieve their related neighbour category images at the rate of choices 5,10,15 and 20 using hybrid algorithm. According to experimental results, the ‘Benign’ category has consumed huge time compared to the other two categories. But, the ‘Malignant’ has consumed less amount of time. |
 |
4.0 Conclusion |
| The concepts of Image mining and Content Based Image Retrieval have been combined in this phase in order to classify the MR images to identify and retrieve tumour formation images. Normally, these two concepts are in distinct areas. But, for this research, a platform is created to combine these two methodologies in an effective manner. The hybridized algorithm discussed in the paper (Kannan, A et al, 2013) has been adapted here in addition to NN method to design a CBIR system to assist medical staff to take relevant action for the tumour conditioned patients earlier. Over 150 patients’ MR images have been taken from DICOM and classified for the research. All the images taken are supervised images since those are already concluded as tumour affected images and labelled each one. |
| The central objective of this paper is to diagnose the given query MR image in a better manner with high accuracy rate and low error rate compared to earlier approaches. The CBIR designed in this phase is to act as a medical decision support system to support the physicians in much more level to the welfare of the patients. This research work will be very much useful to the physicians to take immediate remedial actions for patients without consultation of expert and also the time consumption for taking remedial actions will be consistently low. |
References |
|