e-ISSN: 2319-9849
e-ISSN: 2319-9849
Industrial Unit, Department of Chemistry, Faculty of Science, Helwan University, Cairo, Egypt
Received Date: 22/03/2018 Accepted Date: 30/03/2018 Published Date: 06/04/2018
Visit for more related articles at Research & Reviews: Journal of Chemistry
Semicarbazide (SC) and thiosemicarbazide (TSC) derivatives were allowed to react with salicylaldehyde (SA), 2-acetylpyridine (AP) and 2-acetylthiophene (AT). The synthesized ligands are reacted with some transition metals like Ni and Cu with molar ratios 2:1 and the prepared complexes are characterized by spectroscopic methods. The expected structures for the prepared complexes, total energy, dipole moment and dihedral angles were theoretically studied and fully optimized using Gaussian 09w.
Semicarbazide, Thiosemicarbazide, Biological activity, Transition metal complexes
The modern study of coordination compounds begins with two famous men, Alferd Werner and Sophus Mads Joargensen. Alferd Werner in 1893, proposed the coordination theory. For this pioneering work Alferd Werner received the Nobel Prize in 1913. In fact he was the founder of modern coordination chemistry who postulated the first successful theory, known as “Werner’s Coordination Theory”, to study the formation, properties, characteristics and geometric stereochemistry of coordination compounds [1]. It was proposed that a chemical bond required the sharing lone pair of electrons lead to the idea that a neutral molecule with an electron pair (Lewis–base) can donate these electron to vacant orbitals on metal ion or other electron acceptor (Lewis acid). Although the electron pair donor concept of Lewis is still useful for many Lewis acid base interactions for complex formation. The detailed and more modern concepts to explain formation of coordination bonds, the associated bond properties, structures, stabilities, geometrical shape and the molecular properties as a whole are more conveniently and successfully considered in terms of modern bonding theories the VBT, the CFT, the LFT and the MOT. Metals play a very important role in an immense number of extensively differing biological processes. Some of these processes are quite specific in their metal ion requirements, in that only certain d-block ions in certain oxidation states can accomplish the necessary catalytic structural requirement [2].
Synthesis of [Cu (CH3COO-)2(SASC)2]H2O (M5) Complex
The ethanolic solution of 0.005 mol copper acetate monohydrate was mixed with ethanolic solution of 0.01 mol of (SASC) ligand and the solution was refluxed for 4h. The completeness of reaction was followed by TLC and the final product was diltered and dried [3].
Synthesis of [NiCl2(APSC)2].6H2O (M8) Complex
An aqueous solution of 0.005 mol nickel chloride hexahydrate was added to an ethanolic solution of 0.01 mol of (APSC) ligand with continuous refluxing for 2h. The completeness of reaction was followed by TLC and the final product was filtered and dried [4].
Synthesis of [NiCl2(ATSC)2].6H2O (M13) Complex
An aqueous solution of 0.005 mol nickel chloride hexahydrate was mixed with hot ethanolic solution of 0.01 mol of (ATSC) ligand with continuous stirring at 90°C for 3h. The completeness of reaction was followed by TLC and the final solution was filtered and dried [5].
Mass Spectrum of M5 Complex
Yield%, 71%; m.p. 173°C. The mass spectrum of the prepared M5 complex didn’t show the molecular ion peak of the complex due to its unstability, the suggested structure is 2A1.HCl+Cu+2 +2(CH3COO)-+H2O, which has a molecular weight 630.9 g/mol (Figure 1). The peak appeared at m/z=106 due to (Ph (OH)-CH). The peak appeared at m/z=301 due to ((Ph (OH)-CH=N…Cu (OAc)2) [6,7].
IR Spectrum of M5 Complex
The IR spectrum of the complex M5 was recorded in KBr Cell. It showed a band at 1589 cm-1 due to C=N (azomethine) which may be shifted to lower wave number due to binding with metal (Figure 2). The bands appeared at 3458 cm-1 due to OH which is coordinated with metal. The band appeared at 474 cm-1 may be due to [O→Cu+2].
The expected structure is:
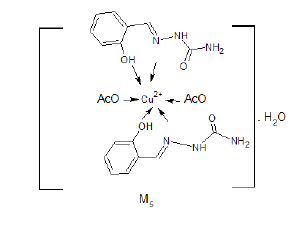
Mass Spectrum of M8 Complex
Yield%, 64%; m.p, 163°C. The mass spectrum of the prepared M8 complex didn’t show the molecular ion peak of the complex due to its unstability, the suggested structure is 2C1.HCl+Ni+2 +2Cl -+6H2O, which has a molecular weight 668.15 g/mol. The peak appeared at m/z=85 due to (C=N-NH-C(O)-NH2) (Figure 3). The peak appeared at m/z=299 due to ((C=N-NH-C(O)-NH2))2--- NiCl2. The peak appeared at m/z=152 due to removal of C2H2 from pyridine ring of C1 ligand.
IR Spectrum of M8 Complex
The IR spectrum of the complex M8 was recorded in KBr Cell. It showed band at 1663 cm-1 due to C=O which may be shifted to lower wave number due to binding with metal also, the band appeared at 1523 cm-1 due to C=N (azomethine group) which may be shifted to lower wave number due to binding with metal (Figure 4). The bands appeared at 3300-3380 cm-1 due to different free NH2 groups. The band appeared at 563 cm-1 may be due to [N→Ni+2] [8,9].
The expected structure is:
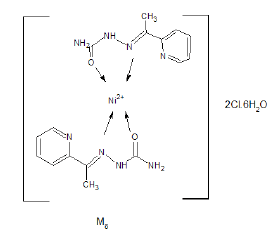
Mass Spectrum of M13 Complex.
Yield%, 52%; m.p, 154°C. The mass spectrum of the prepared M13 complex didn’t show the molecular ion peak of the complex due to its unstability, the suggested structure is 2B1.HCl+Ni+2 +2Cl-+6H2O, which has a molecular weight 677.12 g/mol. The peak appeared at m/z=143 due to (N…NiCl2) (Figure 5). The peak appeared at m/z=338 due to (CH3-C=N-NH-C(O)-NH2 ....NiCl2.6H2O [10,11].
IR Spectrum of M13 Complex
The IR spectrum of the complex M13 was recorded in KBr Cell. It showed band at 1638 cm-1 due to C=O which may be shifted to lower wave number due to binding with metal also, the band appeared at 1433 cm-1 due to C=N (azomethine group) which may be shifted to lower wave number due to binding with metal (Figure 6). The bands appeared at 3350-3470 cm-1 due to different free NH2 groups. The band appeared at 575 cm-1 may be due to [N→Ni+2] [12,13].
The expected structure is:
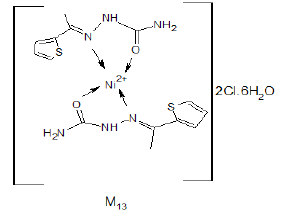
The magnetic properties for the prepared complexes are summarized in Table 1.
Table 2 results show that the structure M8 and M13 have the total energy -3638.27040 and -4247.76657 a.u, which their geometrical structures are tetrahedral structures, which expected to be the most stable structures through a formation of coordination bond with carbonyl and imine groups, Nickel bonded with two chloride ions to give a total coordination number 4 [14].
| Complex | Saturation magnetization (Ms), | Remanence magnetization (Mr), | The coercive field (Hc) | Effective magenetic moment, B.M | No of unpaired electrons | Expected structure |
|---|---|---|---|---|---|---|
| M5 | 0.01673 | 0.00196 | 146.84 | 1.98 | 1 | Octahedral |
| M8 | 0.03105 | 0.00252 | 105.07 | 3.25 | 2 | Tetrahedral |
| M13 | 0.02430 | 0.00220 | 159.47 | 2.94 | 2 | Tetrahedral |
Table 1. Saturation magnetization, remanence magnetization, coercive field and magnetic moment for the prepared complexes.
| Complex | M13 | M8 | ||
|---|---|---|---|---|
| Structure labeling | 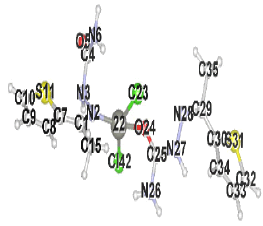 |
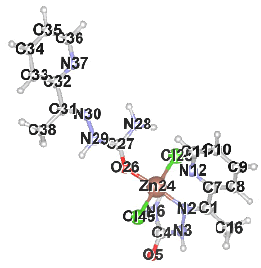 |
||
| Structure | 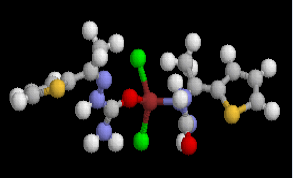 |
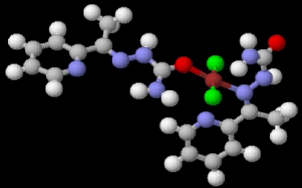 |
||
| Etotal, a.u. | -4247.76657 | -3638.27040 | ||
| Sum of ΔH | -4247.92898 | -3638.47949 | ||
| Sum of ΔG | -4248.02671 | -3638.57416 | ||
| EHOMO, a.u. | -0.21025 | -0.20055 | ||
| ELOMO, a.u. | -0.07596 | -0.09746 | ||
| ΔEg ev | ||||
| Atomic charges | N2 | -0.431 | N2 | -0.420 |
| N3 | -0.503 | N3 | -0.472 | |
| O5 | -0.507 | O5 | -0.488 | |
| N6 | -0.757 | N6 | -0.776 | |
| O24 | -0.516 | N12 | -0.583 | |
| N26 | -0.765 | O26 | -0.617 | |
| N27 | -0.503 | N28 | -0.740 | |
| N28 | -0.215 | N29 | -0.472 | |
| S31 | 0.287 | N30 | -0.270 | |
| --------- | ------------ | N37 | -0.452 | |
| Bond length (R), Å | R(1,2) | 1.305 | R(1,2) | 1.307 |
| R(1,7) | 1.451 | R(1,7) | 1.461 | |
| R(2,3) | 1.415 | R(2,3) | 1.382 | |
| R(3,4) | 1.425 | R(3,4) | 1.441 | |
| R(4,5) | 1.219 | R(4,5) | 1.216 | |
| R(4,6) | 1.366 | R(4,6) | 1.365 | |
| R(7,8) | 1.389 | R(7,8) | 1.394 | |
| R(2,22) | 1.863 | R(7,12) | 1.360 | |
| R(8,9) | 1.413 | R(8,9) | 1.393 | |
| R(22,23) | 2.209 | R(9,10) | 1.3929 | |
| R(22,24) | 1.877 | R(10,11) | 1.3928 | |
| R(24,25) | 1.245 | R(24,25) | 3.0848 | |
| R(25,26) | 1.362 | R(25,26) | 1.8767 | |
| R(25,27) | 1.370 | R(25,27) | 1.2758 | |
| R(27,28) | 1.363 | R(27,28) | 1.3192 | |
| R(28,29) | 1.290 | R(27,29) | 1.3765 | |
| R(29,30) | 1.477 | R(29,30) | 1.3555 | |
| R(29,35) | 1.507 | R(32,37) | 1.347 | |
| R(30,31) | 1.754 | R(30,31) | 1.2888 | |
| R(31,32) | 1.730 | R(31,32) | 1.4911 | |
| R(32,33) | 1.368 | R(32,33) | 1.4036 | |
| R(33,34) | 1.424 | R(33,34) | 1.3943 | |
| Bond angle (A)° | A(2,1,7) | 125.882 | A(2,1,7) | 112.720 |
| A(1,2,3) | 115.845 | A(1,2,3) | 118.098 | |
| A(2,3,4) | 120.621 | A(2,3,4) | 122.768 | |
| A(3,4,5) | 118.515 | A(3,4,5) | 117.587 | |
| A(3,4,6) | 115.524 | A(3,4,6) | 115.965 | |
| A(5,4,6) | 125.609 | A(5,4,6) | 126.360 | |
| A(1,7,8) | 124.010 | A(1,7,8) | 125.269 | |
| A(1,2,22) | 129.800 | A(1,7,12) | 113.323 | |
| A(3,2,22) | 113.947 | A(8,7,12) | 121.404 | |
| A(7,8,9) | 113.897 | A(7,8,9) | 119.025 | |
| A(2,22,23) | 91.718 | A(8,9,10) | 119.058 | |
| A(2,22,24) | 88.532 | A(9,10,11) | 118.980 | |
| A(22,24,25) | 132.927 | A(7,11,12) | 119.443 | |
| A(24,25,26) | 123.670 | A(2,24,25) | 82.452 | |
| A(24,25,27) | 120.536 | A(2,24,26) | 175.478 | |
| A(26,25,27) | 115.718 | A(24,25,26) | 96.778 | |
| A(25,27,28) | 120.062 | A(24,26,27) | 122.320 | |
| A(27,28,29) | 118.953 | A(26,27,28) | 125.621 | |
| A(28,29,30) | 125.360 | A(26,27,29) | 115.590 | |
| A(28,29,35) | 116.105 | A(28,27,29) | 118.699 | |
| A(30,29,35) | 118.534 | A(27,29,30) | 118.981 | |
| A(29,30,31) | 120.816 | A(29,30,31) | 119.599 | |
| A(29,30,34) | 129.140 | A(30,31,32) | 115.775 | |
| A(31,30,34) | 109.996 | A(30,31,32) | 121.277 | |
| A(30,31,32) | 91.894 | A(31,32,37) | 116.778 | |
| A(31,32,33) | 111.847 | A(33,32,37) | 121.936 | |
| A(32,33,34) | 112.651 | A(32,33,34) | 119.113 | |
| ------- | ------- | A(1,2,24) | 116.800 | |
| Dihedral angle (D) ° | D(7,1,2,3) | -1.4548 | D(7,1,2,3) | 177.298 |
| D(2,1,7,8) | -179.546 | D(2,1,7,8) | -177.7415 | |
| D(1,2,3,4) | 113.37 | D(1,2,3,4) | -73.2758 | |
| D(2,3,4,5) | 161.84 | D(2,3,4,5) | -177.2052 | |
| D(2,3,4,6) | -24.5704 | D(2,3,4,6) | 5.9724 | |
| D(7,1,2,22) | -173.6101 | D(7,1,2,2 4 ) | -0.0225 | |
| D(22,2,3,4) | -73.2186 | D( 1 2, 7 , 8 , 9 ) | 0.2634 | |
| D(1,2,22,23) | -73.5074 | D(1,2,2 4 ,2 6 ) | -2.7323 | |
| D(1,2,22,24) | -164.68 | D(1,2,2 4 ,2 5 ) | 77.7595 | |
| D(3,2,22,23) | 114.217 | D(3,2,2 4 ,2 5 ) | -99.3536 | |
| D(3,2,22,24) | 23.0443 | D(3,2,2 4 ,2 6 ) | -179.8454 | |
| D(1,7,8,9) | -179.3754 | D(1,7,8,9) | 179.7115 | |
| D(22,24,25,26) | 6.4254 | D( 1 , 7 , 12 , 11 ) | -179.6059 | |
| D(22,24,25,27) | -170.2927 | D( 8 , 7 , 12 , 11 ) | -0.0965 | |
| D(24,25,27,28) | -5.853 | D(24,2 6 ,27,28) | -40.5313 | |
| D(26,25,27,28) | 177.1784 | D(24,2 6 ,27,2 9 ) | 142.937 | |
| D(25,27,28,29) | -179.4751 | D(2 6 ,2 7 ,2 9 , 30 ) | 179.3258 | |
| D(27,28,29,30) | -3.4089 | D(2 8 ,2 7 ,29,30) | 2.5397 | |
| D(27,28,29,35) | 176.4606 | D(27,2 9 , 30 ,3 1 ) | 169.5779 | |
| D(28,29,30,31) | 137.643 | D(29,30,31,32) | 179.0346 | |
| D(28,29,30,34) | -45.0917 | D(2,1,7,12) | 1.7456 | |
| D(35,29,30,31) | -42.2236 | D(35,36,37,32) | -0.0041 | |
| D(35,29,30,34) | 135.0417 | D(3 4 ,3 5 ,36,37) | 0.3304 | |
| D(29,30,31,32) | 178.1294 | D(3 3 ,3 4 ,3 5 ,36) | -0.406 | |
| D(34,30,31,32) | 0.3862 | D(3 2 ,3 3 ,3 4 ,3 5 ) | 0.1838 | |
| D(29,30,34,33) | -177.7428 | D(3 3 ,3 2 ,3 7 ,3 6 ) | -0.2404 | |
| D(31,30,34,33) | -0.2418 | D(31,3 2 ,3 7 ,3 6 ) | -179.2626 | |
| D(30,31,32,33) | -0.4449 | D(30,31,32,33) | -158.4369 | |
| D(31,32,33,34) | 0.3865 | D(31,32,33,3 7 ) | 20.5921 | |
| D(32,33,34,30) | -0.0897 | D(3 7 ,3 2 ,3 3 ,3 4 ) | 0.1503 | |
Table 2. The total energies, enthalpies, bond angles, dihedral angles for the prepared complexes.
The reaction between semicarbazide, thiosemicarbazide with aldehydes and ketones groups are used for Schiff base derivatives preparation, the prepared ligands have multidentate function groups which increase their ability for binding with transition metals. The study of magnetic moment and theoretical calculations prove the geometrical shape of the prepared complexes.