ISSN ONLINE(2319-8753)PRINT(2347-6710)
ISSN ONLINE(2319-8753)PRINT(2347-6710)
M.P.Kusuma1 , D.Sri Rami Reddy2, Mukund Sharma3
|
| Related article at Pubmed, Scholar Google |
Visit for more related articles at International Journal of Innovative Research in Science, Engineering and Technology
Pectinases are an enzyme group that catalyzes pectin substance degradation through depolymerization (hydrolases and lyases) and deesterification (esterases) reactions. One of the most important and widely used commercial pectinase is polygalacturonase. Pectinolytic enzymes are naturally produced by many organisms like bacteria, fungi, yeasts, nematodes, protozoan and plants. Alkaline pectinases are generally produced by bacteria particularly species of Bacillus, but are also made by some filamentous fungi and yeasts (1) and acidic pectinases are produced by fungi. A total of 66 bacterial colonies were isolated from six sampling sites. Out of 66 isolates 8 isolates showed polygalacturonase actity and C4 isolate showed maximum actity. The potential new isolate on polygalacturonase production was tested against three polygalacturonase producing microorganism. Comparately isolate C4 showed maximum enzyme actity hence it was chosen for in depth study. The isolate C4 was characterized and identified as Bacillus subtilis C4 by biochemical studies and 16 rDNA analysis. The new Bacillus subtilis was used for polygalacturonase production under submerged ferementation. Growth profile of the above isolate was studied. The results revealed that the organism was showing maximum log phase till 24hr period after which the growth gradually reduced and remained stable till 72h indicating stationary phase at 400C and pH 11.0
INTRODUCTION |
| A large number of industrial processes in the areas of industrial, environmental and food biotechnology utilize enzymes. Today, enzymes for a major subject of research and because of their specificity of action are of great value as analytical reagents (2).The increasing energy demands have focused worldwide attention on utilization of renewable resources, particularly agricultural and forest residues, the major components of which are cellulose, starch, lignin, xylan and pectin. Pectin was first isolated and described by Henri Braconnot (1825). Pectin is jelly like matrix structural polysaccharide found in primary cell wall and middle lamellae of fruits and vegetables (Torres et al, 2006). Three major pectic polysaccharide groups were recognized all, containing D-galacturonic acid to a greater or lesser extent namely Homogalacturonan, Rhamnogalacturonan I and Rhamnogalacturonan II. Pectic substances tend to form gel structure when portions of homogalacturonan are cross linked forming a three dimensional crystalline network at which water and solutes are trapped. These polysaccharides have been used as a bioactive food ingredient and as detoxifying agent.Pectinases are an enzyme group that catalyzes pectin substance degradation through depolymerization (hydrolases and lyases) and deesterification (esterases) reactions. One of the most important and widely used commercial pectinase is polygalacturonase. Pectinolytic enzymes are naturally produced by many organisms like bacteria, fungi, yeasts, nematodes, protozoan and plants. Alkaline pectinases are generally produced by bacteria particularly species of Bacillus, but are also made by some filamentous fungi and yeasts (3) and acidic pectinases are produced by fungi. Pectinase production occupies about 10% of the overall manufacturing of enzyme preparations. These enzymes are widely used in the food industry in the production of juices, fruit drinks and wines (4). Submerged fermentation and solid state fermentation have been successfully used in pectinase production by fungi (5) and bacteria (6). They may be used in the pretreatment of waste water from vegetable food processing that contains pectin residues, the processing of textile fibers such as flax, jute and hemp, coffee and tea fermentation, vegetable-oil extraction and the treatment of paper pulp(7).Realizing the importance of pectinase, the present investigation was aimed with the Screening of suitable microorganism and substrate for the production of polygalacturonase under SMF and optimization of microbial parameter for the production of polygalacturonase. |
MATERIALS AND METHODS |
| II-1) Screening and isolation of polygalacturonase producing bacteria |
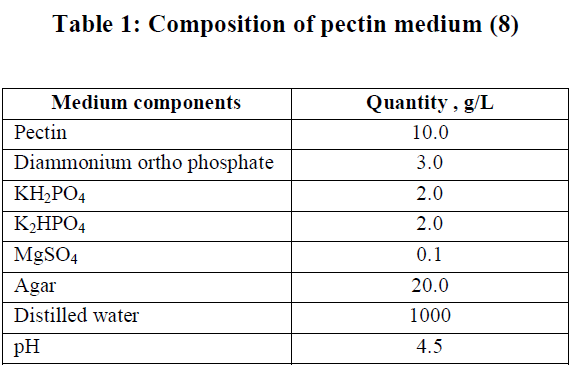
| Derse habitats of different areas were selected for isolation of bacterial strains with a view to isolate acte producer of polygalacturonase. Four samples were taken up to depth of 20cm after removing approximately 3cm of the soil surface. All the samples were collected in the sterile screw capped tubes and care was taken to see that the points of collection had as widely varying characteristics as possible with regard to organic matter, moisture content, particle size and colour of soil.1gm each of the soil samples was taken into 100mL of sterile distilled water and the suspension was kept in rotary shaker for 30 min and kept aside to settle suspending matter. One mL of the suspension was serially diluted with sterile water. Pure cultures were isolated and screened for extracellular polygalacturonase using agar media containing pectin as a substrate (Table 1). The plates were incubated in an inverted position for 48h at 370C. Three replicates were prepared for each sample. |
 |
| Cultures which were having potential polygalacturonase actity were sub-cultured on the slant and maintained for identification and enzyme production. |
| II-2-PLATE ASSAY OF DEPOLYMERIZED PECTIN (9) |
| Pure culture was inoculated into the pectin agar medium and incubated at 37ºC for 48h. After the colonies reached around 3 mm, iodine-potassium iodide solution (1.0g iodine, 5.0g potassium iodide and 330mL H2O) was added to detect clear zones. Of the 66 isolated tested for polygalacturonase production, 8 isolates have shown polygalacturonase actity and the C4 isolate exhibited high polygalacturonase actity. (Tables 3.1(a and b). The promising C4 isolate was compared with three other cultures procured from MTCC, Chandigarh (Table 2.2). The isolate was checked for its actity in the fermentation medium (Table 2.3). Further it was subjected to detailed morphological and biochemical studies. |
| II-3)-IDENTIFICATION OF THE ISOLATE C4 BASED ON MORPHOLOGICAL AND BIOCHEMICAL CHARACTERISTICS |
| Various morphological and biochemical characteristic tests like methyl red test, Voges Proskauer, Casein hydrolysis, Citrate utilization, Nitrate test, Indole, Arginine, Lysine, Ornithin, Gelatin hydrolysis, Starch hydrolysis, Esculin hydrolysis, Catalase test, Oxidase test and growth on Mc Conkey were performed . |
| II-4)-16S RDNA ANALYSIS |
| II-4-1) ISOLATION OF GENOMIC DNA FROM BACTERIA ( 10) |
| 5 mL of bacterial culture was grown to saturation and centrifuged at 10,000 rpm for 2 min. The pellet was resuspended in 567 μl TE buffer by repeated pipetting. 30 μl of 20% SDS, 5 μl of proteinase K (25 mg/mL) and 5 μl of RNase (10 mg/mL) and 30 μl lysozyme (100 mg/mL) was added to the suspension and incubated at 37 °C for 1 h. 100 μl of 5 M NaCl, 80 μl CTAB (cetyltrimethylammonium bromide)/NaCl solution was added to it and incubated at 65 °C for 30 min. Equal volumes of chloroform/isoamyl alcohol was added, mixed well and centrifuged for 5 min at room temperature. The upper aqueous phase was transferred to fresh eppendorf without disturbing the interface and equal volume of phenol/chloroform/isoamyl alcohol (25:24:1) was added to it, mixed by inversion and centrifuged for 10 min at room temperature. To the supernatant, 0.6mL isopropanol was added, mixed well and again centrifuged at 15,000 rpm for 15 min at 4 °C. The pellet was washed with 70% ethanol and dried at 4 °C. The dried pellet (using Speed Vac/ laminar air flow) was resuspended in sterile distilled water. The concentration of the resuspended DNA was quantitated spectrophotometrically and quality was checked by agarose gel electrophoresis. |
| II-4-2) 16S RDNA AMPLIFICATION |
| The above 16s rDNA was amplified by subjecting the above obtained DNA to PCR using appropriate markers and subjected to agarose gel electrophoresis. The obtained DNA fragment was excised with a clean, sharp scalpel. The gel was weighed in a in a colorless tube. 3 volumes of buffer QG was added to 1 volume of gel. Incubated at 50 ºC for 10 min (or until the gel slice has completely dissolved and the color of the mixture is yellow). The adsorption of DNA to the membrane is efficient only at pH ≤7.5. Buffer QG contains a pH indicator, which is yellow at pH ≤7.5 and orange or violet at higher pH, allowing easy determination of the optimal pH for DNA binding. To bind DNA, apply the sample to the MiniElute column, and centrifuge for 1 min. To elute DNA, 10 μl of autoclaved double distilled water or buffer EB (10 mM Tris·Cl, pH 8.5) was added at the center of the membrane, let the column stand for 1 min, and centrifuged for 1 min. |
| II-4-3) 16S RRNA GENE SEQUENCING |
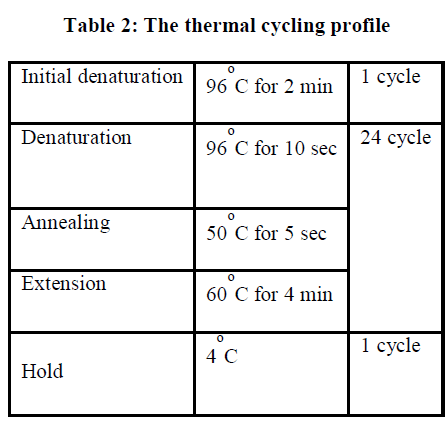
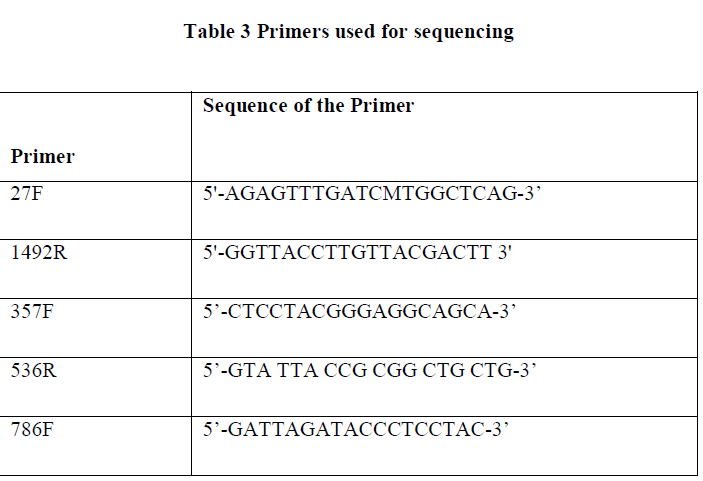
| Full length sequence of 16S rRNA gene can be achieved by using three different internal primers with three different sequencing PCRs. For each reaction the following reagents are added. |
| II-4-4) REACTION VOLUME ADDED |
| Terminator Ready Reaction Mix 4 μl |
| PCR product x μl |
| Primer (3.2ng/ μl) 1 μl |
| Double distilled water added to final Vol. |
| Total volume 10 μl |
 |
| II-4-5) POST REACTION CLEAN UP AND RESUSPENSION (REMOVAL OF UNINCORPORATED DYE TERMINATORS FROM THE SEQUENCING REACTION |
| 2 μl of 125 mM EDTA was added to each reaction tube containing 10μl of reaction mixture and the contents are mixed. 50μl of 95% ethanol was added to each tube and incubated at room temperature for 15 min. Spinned at 12,000 rpm for 20 min at room temperature and to each tube 250μl of 70% ethanol (at RT) added. Repeat spinning and decant and finally wash with 70% ethanol wash. Aspirate or decant the supernatant and air dry the pellet for 30-45 min. Dried pellet can be stored at -20 ºC for several weeks. |
| II-4-6) LOADING SAMPLE FOR SEQUENCING |
| 10-12μl of Hi Di foramide was added to the dried pellet. Heated at 95 ºC for 30 min and immediately chilled on ice. Briefly the tube was spinned and the content was transferred to 96 well reaction plate and submit it for sequencing. |
| II-4-7) Sequencing |
| Sequencing was done in autosequencer based on Sanger’s method of sequencing. Chromatograms obtained were analysed on finch TV. Assembly was done on DNA star. |
 |
| II-5) SCANNING ELECTRON MICROSCOPIC OBSERVATION OF THE ISOLATE C4 |
| Samples were fixed in 2.5% gluteraldehyde in 0.1M phosphate buffer(pH 7.2) for 24h at 40C and post fixed 2% in aqueous osmium tetra oxide for 4 h. Dehydrated in series of graded alcohols and dried to critical point drying with CPD unit. The processed samples were mounted over the stubs with double-sided carbon conductity tape, and a thin layer of gold coat over the samples were done by using an automated sputter coater (Model – JEOL JFC-1600) for 3 minutes and scanned under Scanning Electron Microscope (SEM – Model: JOEL –JSM 5600) at required magnifications. |
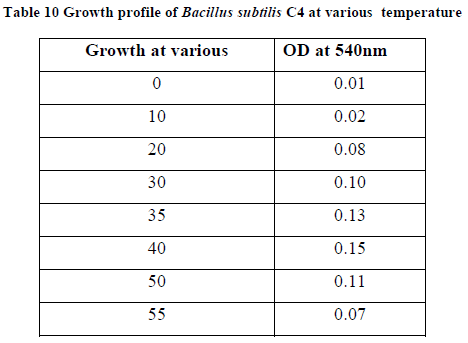
| II-6) GROWTH PROFILE OF BACILLUS SUBTILIS C4 AT VARIOUS TEMPERATURES |
| Overnight slant grown culture suspension was taken into each sterile nutrient broth. 5ml of the sterile nutrient broth was taken in another tube to be used as a blank .The broths were incubated at various temperatures ranging from 00C - 550C. After incubation, 0.5ml of formaldehyde was added to protect it from contaminating bacterial growth. O.D was taken at 540nm against blank. |
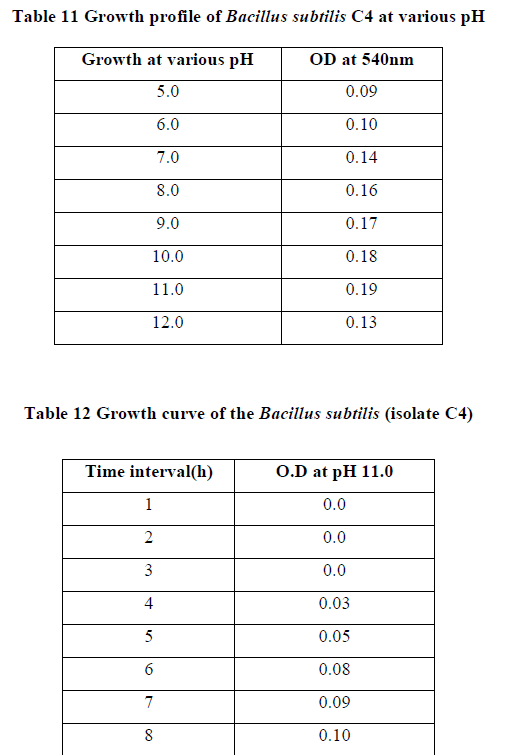
| II-7) GROWTH PROFILE OF BACILLUS SUBTILIS C4 AT VARIOUS PH |
| Overnight slant grown culture suspension was taken into each sterile nutrient broth. pH of each of the medium was adjusted in the range of 5.0-12.0 and incubated. After incubation, 0.5ml of formaldehyde was added to protect it from contaminating bacterial growth. O.D was taken at 540nm against blank. |
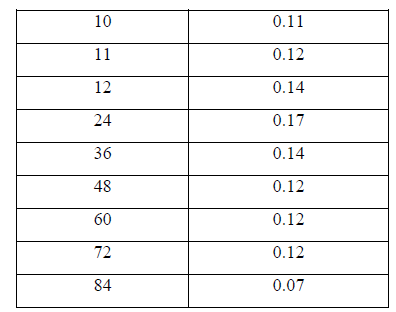
| II-8) GROWTH CURVE OF BACILLUS SUBTILIS C4 |
| Overnight slant grown culture suspension was taken into each sterile nutrient broth. 5ml of the sterile nutrient broth was taken in another tube to be used as a blank .The broths were incubated for various time intervals. After incubation, 0.5ml of formaldehyde was added to protect it from contaminating bacterial growth. O.D was taken at 540nm against blank. |
| II-9) POLYGALACTURONASE PRODUCTION BY SUBMERGED FERMENTATION |
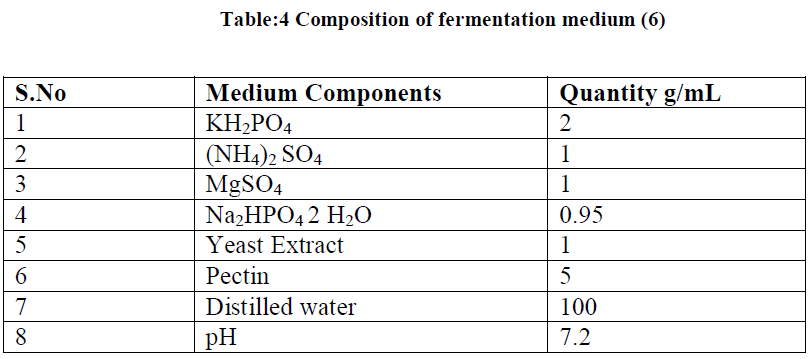
| The fermentation medium was prepared by using following ingredients (Table 2.4). The flasks were sterilized at 1210C at 15lb/sq inch pressure. The sterilized medium was inoculated with 1mL of 12h culture of Bacillus subtilis maintaining a pH of 7.2. After 2days of incubation the medium was centrifuged at 5000rpm for 15min and the supernatant was taken for the assay of amount of galacturonic acid produced. |
 |
| II-10) POLYGALACTURONASE ASSAY (11) |
| Polygalacturonase actity was assayed by the colorimetric method of Miller (1959). 0.5mL of supernatant was incubated with 1.0% of pectin at 400C for 10min under static conditions. After adding 3mL of di nitro salicylic acid, the mixture was boiled for 15min and finally diluted to 5mL with distilled water. The absorbance was measured at 540nm. |
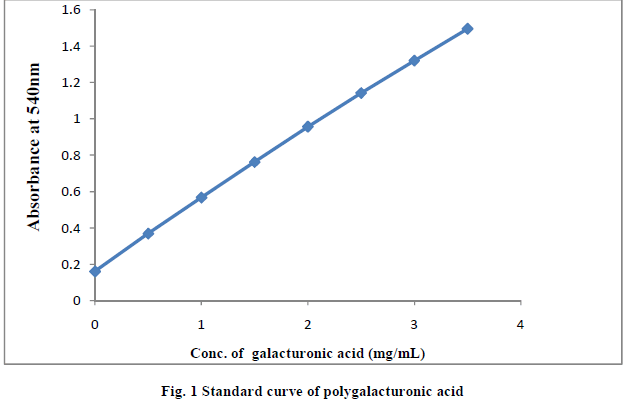
| II-11) STANDARD CURVE PROTRACTION (12) |
| 2mL galacturonic acid solution of 10mg/mL was diluted to 100mL to obtain d-galacturonic solution of 0.25mg/mL and then it was treated with DNS reagent. The mixed solution was boiled for 30min. After cooling, solution was mixed with 10mL of distilled water in tubes. Finally, absorbency of the solution at 540 nm was measured |
 |
| II-12) EFFECT OF MICROBIAL PARAMETERS |
| Microbial parameters like inoculum age, inoculum volume and agitation speed The strategy adopted for standardization of fermentation parameters was to evaluate the effect of an indidual parameter and incorporate it at standard level before standardizing the next parameter. |
| II-13) EFFECT OF INOCULUM VOLUME |
| To determine the effect of inoculum volume on the production of polygalacturonase, each flask was sterilized, inoculated with 0.8×102CFU/mL different inoculum volumes of 0.75,1.0,2.5,5.0,7.5,10,12.5 and 15mL of 12h inoculum and incubated. |
| II-14) EFFECT OF INOCULUM AGE |
| In order to determine the optimum inoculum age, each flask was inoculated with inoculum of varying inoculum age ranging from 12h to 48h with 12h interval and assayed for enzyme actity. |
| II-15) EFFECT OF AGITATION SPEED |
| In order to determine the optimum agitation speed, each flask with fermentate medium was inoculated and subjected to various rpm viz. 120,150,180 and 210, incubated and assayed for the presence of enzyme. |
EXPERIMENTAL |
| III.1) ISOLATION OF POLYGALACTURONASE PRODUCING BACTERIA |
| A total of 66 colonies were isolated from four soil samples, on the basis of macroscopic characters and microscopic observations, eliminating those that appeared close to each other. All 66 isolated colonies were tested for their polygalacturonase potential using plate assay. Of the 66 isolated colonies tested, only eight bacterial strains shown the polygalacturonase potential. Of the 8 posite isolates, C4 was showing relately high actity. (Table 5 a and b) |
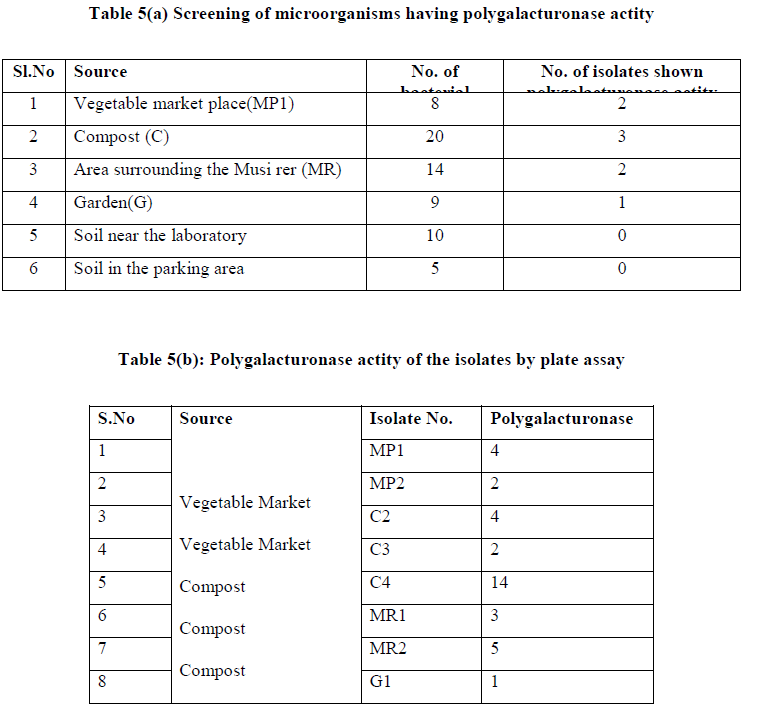 |
| III.2) SCREENING OF MICROORGANISMS FOR HIGH POLYGALACTURONASE PRODUCTION |
| Plate assay was adopted for screening the polygalacturonase (Chesson and Codner (2008)). Pure culture was inoculated into the pectin agar medium and incubated at 37ºC for 48h. After the colonies reached around 3 mm, iodinepotassium iodide solution (1.0g iodine, 5.0g potassium iodide and 330mL H2O) was added to detect clear zones. Four cultures were used for the assay, out of which Streptomyces lydicus (MTCC 4067), Bacillus pumilus (MTCC 1456), Aspergillus niger (MTCC 281) were procured from MTCC, Chandigarh and isolate C4 isolated from soil. Bacillus pumilus (MTCC 1456), Aspergillus niger (MTCC 281) and isolate C4 shown maximum clear zone of 14mm and Streptomyces lydicus (MTCC 4067) has shown only 10mm zone (Table 6 and Fig.2). Polygalacturonase production was also studied in the fermentation medium by all the cultures and their actities were measured. Though in the plate assay, the isolate C4, Bacillus pumilus and Aspergillus niger had shown similar zone of colour change, but in quantitate assay, the isolate C4 has shown more polygalacturonase actity. The promising isolate C4 was identified based on microscopic and biochemical actities. Hence the isolated strain was chosen for detailed study (Table 7). |
| Table 6 Comparison of polygalacturonase production of the new isolate with procured species |
 |
 |
| Table 7 Screening of microorganisms for high polygalacturonase production by SMF |
 |
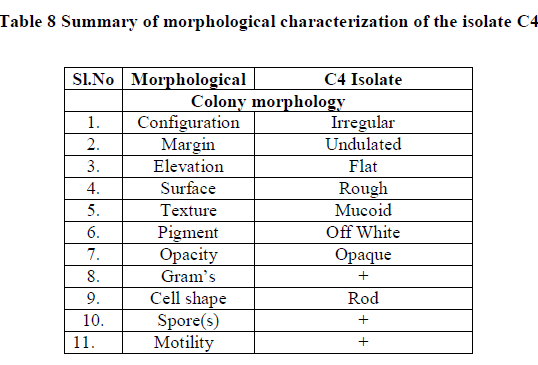
| III.4) MORPHOLOGICAL AND BIOCHEMICAL CHARACTERIZATION OF THE ISOLATE |
| The isolate was proceeded for microscopic and biochemical characterization. |
| Summary of morphological and biochemical characterization of the isolate was gen in the Table 8 & 9 |
 |
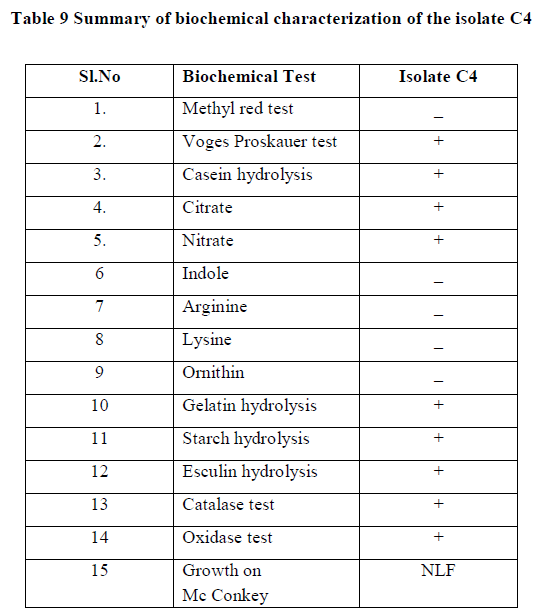 |
| +: Posite ; âÃâ¬Ãâ : Negate ; NLF : Non Lactose fermenting; |
| Correlating the above results with Bergey’s manual of determinate bacteriology infers that, isolate C4 belonged to Bacillus species. 16s r DNA analysis was carried and the isolate C4 was found to be Bacillus subtilis. |
| III.5) ELECTRON MICROSCOPIC OBSERVATION OF BACILLUS SUBTILISC4 |
| Scanning electron microscopic observation of the isolate was carried out and it was revealed that the rods were in the form of diplobacilli. (Fig.3) |
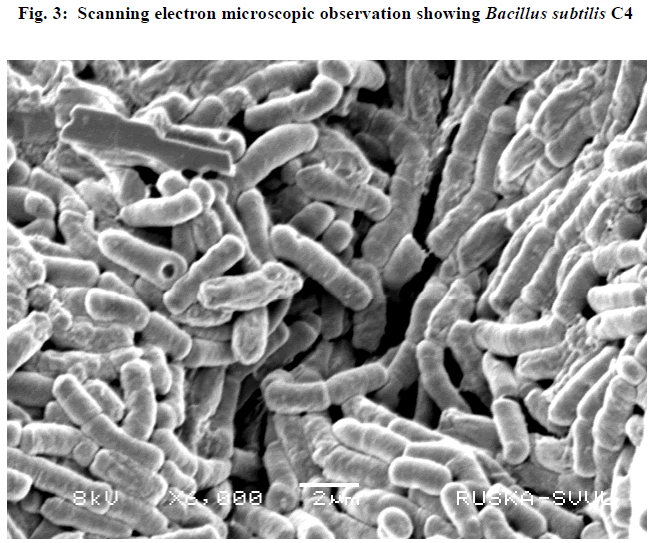 |
| III.6) GROWTH PROFILE OF BACILLUS SUBTILIS (C4) AT VARIOUS TEMPERATURES AND PH |
| Growth profile of the isolate C4 was studied at various temperatures ranging from 00C - 550C. The growth of the bacteria increased gradually with increasing temperatures and finally at 400C maximum growth rate was observed. (Table 10). Further increase in temperature decreased the growth rate due to variety of factors like such as inactation or denaturation of proteins, instability or RNA inhibition (13).The growth was studied at various pH ranging from 5.0-12.0. The growth proceeds rapidly and maximum growth was observed at pH 11.0, and thereafter the growth rate declined (Table 11). |
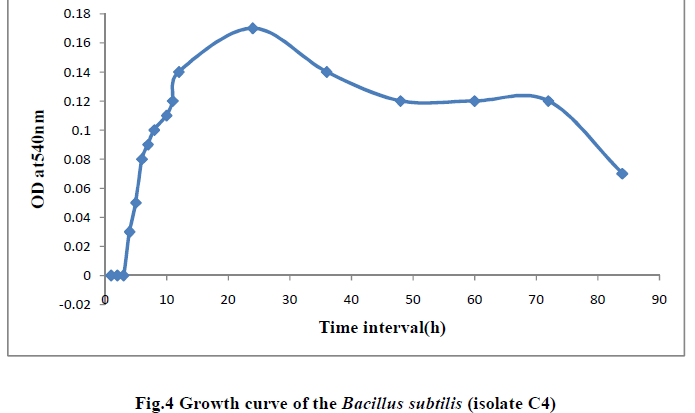
| III.7) GROWTH CURVE OF BACILLUS SUBTILIS C4 AT TEMPERATURE 400C AND PH 11.0 |
| The growth curve of Bacillus subtilis C4 was studied at pH11.0 and temperature 400 C. Initially there was no growth indicating lag phase and the growth started from 4h, increased gradually showed maximum log phase till 24hr period. Thereafter the growth reduced and from 48th hr the growth rate remained stable indicating stationary phase and at 84h the growth rate declined indicating death phase (Table 12 and Fig.4). |
 |
 |
 |
 |
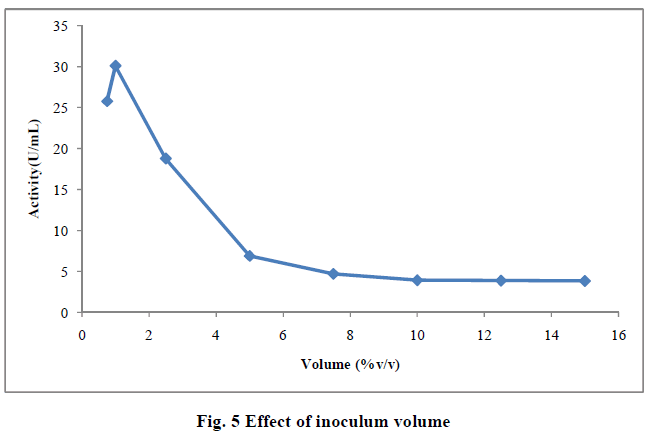
| III.8) EFFECT OF INOCULUM VOLUME |
| The optimum inoculum volume needed to produce the highest yield of enzyme was 1.0mL (30.10U/mL).Optimum inoculum size of 1mL was able to provide enough biomass with optimal length of log phase leading to high levels of enzyme production. Further increase in inoculum decreased the enzyme production. This condition could be due to competition for the nutrients among the bacterial cells. Adequate nutrient supply could be the reason of higher enzyme production with optimum inoculum volume (14). When the inoculum volume was increased more than 1mL, the actity decreased (Fig. 3.4). This may be due to the amount of pectin which was limited in 250mL flask and any increasing inoculum volume does not produce any increase in the yield of polygalacturonase (15) . High inoculum levels are inhibitory in nature and high inoculum density leads to population overcrowding, high nutrient competition and rapid exhaustion of nutrients. The lower inoculum density, may ge insufficient biomass causing reducing product formation (16). Similar results were reported by Reda et al.,(2008) when polygalacturonase was produced by solid state fermentation using Bacillus firmus -I-10104. Maximum pectinolytic actity with inoculum volume of 3% inoculum (6X10 6 spores) and 3 day old spores in submerged fermentation of pectinolytic enzymes by Aspergillus niger (17). |
 |
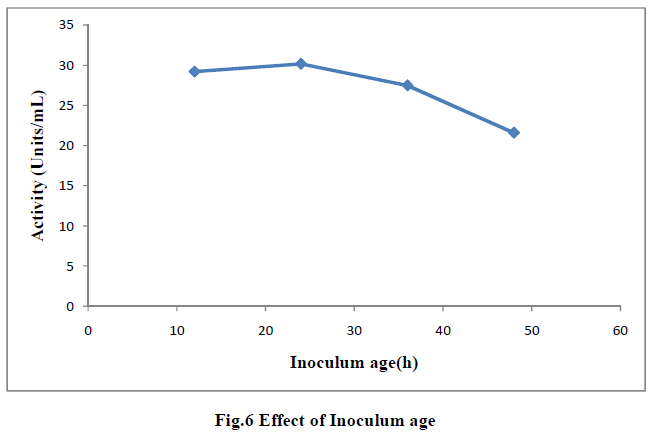
| III.9) EFFECT OF INOCULUM AGE |
| Effect of inoculum age was tested by inoculating the fermentate medium with 1mL inoculum of varying age 12h, 24h, 36h and 48h and the filtered broth was assayed for enzyme actity. Maximum actity(30.17U/mL) was observed when 24h inoculum was used. Thereafter the actity decreased which might be due to the fact that the younger cultures being nonsporulating, entered the growth phase very soon, while the older culture produced spores which took longer time to germinate, and therefore, the enzyme production was low (18) (Fig 6). It was reported that till inoculum age of 16h there was rapid increase in enzyme production and thereafter it became constant when Bacillus sphericus MTCC(7542) was used (19). |
 |
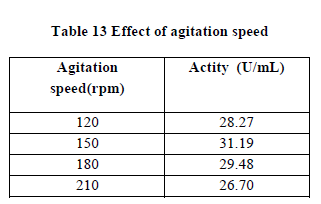
| III.10) EFFECT OF AGITATION SPEED |
| The effect of agitation speed was studied by subjecting the fermentation medium to various rpm viz. 120,150,170 and 200. The optimum agitation speed required to produce greater polygalacturonase production was 150rpm(30.19U/mL)(Table 13). Speed greater or lesser than this produced less polygalacturonic acid. At this speed sufficient aeration must have been present which will provide sufficient dissolved oxygen to the medium (20). |
| It was observed that increase in mixing help the microbial synthesis of enzyme which consequently increases the assimilation of sugars(21). The cell growth started to decline after achieving the maximal production. This condition could be due to shear forces dered from higher agitation speed where collision among the cells occurred and damaged them (14). At lower agitation speed, the inadequate mixing of the broth towards the later stages of growth affected the enzyme synthesis, while the drastic dropped in enzyme actity at higher agitation speeds was due to shearing effect on the cells. Changes in morphology of microorganisms caused by agitation speed also influenced enzyme production and growth of the microorganisms (22). |
 |
CONCLUSION |
| Microbial pectinases have tremendous potential in commercial applications and acidophilic pectinases have extense application in extraction and clarification of juice and wine. However work on the utilization of alkaline pectinases remains underdeveloped as only few reports are available on application of these enzyme Alkaline pectinases have been used in several conventional industrial processes, such as textile and plant fiber processing, coffee and tea fermentation, oil extraction, and treatment of industrial waste water containing pectinacious material. With increased understanding and knowledge of the mechanism of pectin-degrading microorganisms and their enzymes, alkaline pectinases have made their way into several other biotechnological processes such as purification of plant viruses and paper making (23),most of which, despite sound interesting, have yet to be commercialized. The present study suggests that the new soil isolate Bacillus subtilis C4 is a good producer of polygalacturonase.. Considering the process advantages of thermostable enzyme and wide applications of alkaline polygalacturonase, the new soil isolate Bacillus subtilis C4 is a potential candidate for commercial exploitations. |
References |
|