ISSN ONLINE(2319-8753)PRINT(2347-6710)
ISSN ONLINE(2319-8753)PRINT(2347-6710)
Aloo Becky Nancy2, Mulei Josephine1, Mwamburi Aluoch Lizzy3
|
| Related article at Pubmed, Scholar Google |
Visit for more related articles at International Journal of Innovative Research in Science, Engineering and Technology
Slow sand filtration (SSF) is a simple technique for drinking water purification and may be adapted for wastewater stabilization but only a few studies have been done towards this direction. This study aimed at the evaluation of the effect of sand bed depth on SSF performance by comparing data from sewage effluent filtered through sand beds of varying depths (0.5, 0.7 and 1.0) m. Removal efficiency was determined in terms of selected bacteriological indicators such as coliforms, faecal Streptococcus, total bacterial counts and physicochemical parameters such as biochemical oxygen demand, conductivity, pH, Total Suspended Solids and nutrients. Results of this study indicated significant differences in levels of all physicochemical parameters except phosphates in effluents filtered through the three sand bed depths whereby removal efficiency of these parameters was found to be better at 0.7 m. However, removal of bacteria was not significantly affected by sand bed depth.
Keywords |
| Slow sand filtration, wastewater stabilization, bacteria, physicochemical parameters, sand bed depth |
I.INTRODUCTION |
| The world is faced with problems related to management of wastewater due to extensive industrialization and increasing population [1]. Wastewater treatment plants that are aimed at reducing the pollutant load on the environment in most cases release effluents that are still high in Biochemical Oxygen Demand (BOD5), nutrients (N and P) and bacterial load thus posing danger to the receiving environment [2, 3]. Health problems and diseases are often caused by discharging untreated or inadequately treated effluent into waterways. Many infectious diseases are associated with faecally contaminated water and are a major cause of mortality worldwide [4, 5).The release of raw or partially treated effluent into water bodies may also lead to other problems such as fish kills and algal blooms resulting from the high organic contents in the wastewater [6, 7]. |
| Slow sand filtration is a simple technology that can be used to reduce the pollutant load of wastewater [8, 9]. However, little work has been done on the application of Slow Sand Filters (SSFs) in wastewater quality improvement [10]. The process is passive and the effectiveness of the filters is dependent upon the development of a biofilm attached to sand grains called the hypogeal [11]. With increased volumes of treated water being targeted for reuse, there is need to develop reliable methods to mitigate the health risks that can be caused by microorganisms in water. |
| In this study, the effluent from University of Eldoret (UoE) sewage treatment plant was sampled and analysed using standard methods to determine its quality and SSFs of three different sand bed depths were used to filter it thereafter. |
| The objective of this study was to determine levels of physicochemical parameters, bacterial counts and selected indicator bacteria in unfiltered and filtered effluents and to evaluate the effect of sand bed depth on filter performance. |
II. LITERATURE SURVEY |
| Slow sand filtration is a water purification process in which water is passed through a porous bed of sand that contain a biological film that traps and metabolizes the organic compounds in water (Rooklidge et al., 2009). Studies of the effect of sand bed depth on filter performance have also been done by Bellamy et al., (1985), Williams (1987) and Muhammad (1996). Burgoon et al., (1991) studied total phosphate removal in sand beds of different depths and Elliot et al., (2008) studied tertiary wastewater treatment of secondary sewage effluents using laboratory scale models of sand filters. |
III. MATERIALS AND METHODS |
STUDY AREA, DESIGN AND OPERATION OF FILTERS |
| The study was carried out at the (UoE)sewage treatment plant, Uasin Gishu County, Kenya. Samples of filtered and unfiltered effluents were collected on a monthly basis for six consecutive months (June to December 2012). The samples were analysed for bacteriological and physicochemical parameters before introducing into assembled SSFs at the University Biotechnology laboratory.Nine experimental filters were assembled using plastic (PVC) pipes sealed at the bottom and fitted with stainless steel valves. Coarse gravel was used to make the under drain medium of about 0.5 m in each of the filters to hold the filter medium. The effective sand grain size of 1.05 mm was prepared using a standard sieve. The gravel and sand for the filters were washed and sterilized in the oven at 105oC overnight before placing into the plastic pipes. Sand bed depth was0.5 m in the first three sets of filters, 0.7 m in the next three sets and 1.0 m in the last set. Five liters of sampled effluent were introduced into each of the filters and samples were collected after 48 hours for bacteriological and physicochemical analyses. |
BACTERIOLOGICAL ANALYSES |
| Samples were collected and transported in sterilized containers to the laboratory at 4oC and initial analyses were done within 24 hours of sampling. Effluent samples were analysed following standard plating techniques for enumeration of three bacterial types and for total bacterial counts. Positive isolates were selected based on different morphologies on selective media at suitable temperatures of incubation and identified using Gram staining properties and biochemical tests. Standard plating techniques [12] were used to isolate and enumerate Total Coliforms (TC), Faecal Coliforms (FC) and Faecal Streptococcus (FS) and Total Bacterial Counts (TBC) on MacConkey (MAC), Bile Aesculin (BA) and Plate count agars (PCA) and incubated at 37°C for 48 hours. The pink or red colonies growing on MAC agar plates were tested for identification of members of the Enterobacteriaceae family while those growing on BA agar plates were used for identification of FS. The colonies on the BA and MAC agar plates were sub-cultured on Nutrient Agar (NA) and incubated at 37oC for 48 hours to get pure cultures for further identification procedures. Gram staining and microscopy were used to determine the Gram stain reactions and morphologies of bacterial isolates. Gram negative rods gave the initial identification FC and TC bacteria while Gram positive cocci that appeared in chains gave the initial identification of FS.Gram positive cocci bacteria isolated previously on BA agar were subjected to the catalase test using the tube method (South Bend Medical Foundation, 2010) to identify the FS. Four to five drops of 3% hydrogen peroxide were placed into a sterile test tube and a small amount of the pure culture of the test organism added using a sterile wooden applicator stick. The test tube was placed against a dark background and observed for immediate effervescence at the end of the wooden applicator stick [14]. Negative reactions were indicative of the presence of the catalase negative Gram-positive cocci FS [15]. |
| The TBC were determined using standard spread plate technique [12] on PCA and incubated at 35oC for 48 hours. The number of colonies were counted using a Quebec colony counter (Gallenkamp; England) and reported as viable cells. The CFU/ml of samples was calculated by multiplying the counted number of colonies by the dilution used and dividing by total volume plated. |
PHYSICOCHEMICAL ANALYSES |
| Conductivity, temperature and pH of effluent samples were measured in situ using a JENWAY 3405 Electrochemical Analyser and a HACH thermometer and pH meter respectively. Total Suspended solids (TSS) was determined by filtering sample through pre- weighed glass fibre filter pads and drying them overnight at 105°C in the oven to remove any remaining water. The pads were weighed on a weighing balance (Kern & Sohn GMBH, D-72336, Balingen, Germany) and the increase in weight of the filter pads represented the TSS of the effluent samples in mg/l, (APHA, 2005). The amino acid, Diazotization and Cadmium Reduction methods were used to measure phosphates (PO4 -3), nitrites (NO2 -) and nitrates (NO3 -) in water samples using the HACH colorimeter (DR/820).Measurement of Dissolved Oxygen (DO) was carried out using a HANNA DO meter (HI 9143). Determination of BOD5 was done by getting the difference in DO between day 1 of sampling and day 5 (FAO, 2007). |
STATISTICAL ANALYSES |
| Statistical analyses were performed using Statistical Package for Social Sciences (SPSS 15.0; SPSS Inc., IL. USA).Removal efficiencies for physicochemical and bacteriological parameters in the filters were assessed by ANOVA using the general treatment structure in randomized blocks. This was done in order to determine if there were significant differences between influent and effluent parameters. The replications were treated as the blocks and the response parameters were the bacteriological and the physicochemical parameters of the filtered and unfiltered effluent samples. Duncan’s Multiple Range Test (DMRT) was used to separate means that were significantly different. All statistical analyses were done at 95% level of confidence (p < 0.05). |
IV. RESULTS |
ISOLATION AND IDENTIFICATION OF TC, FC AND FS, IN EFFLUENTS |
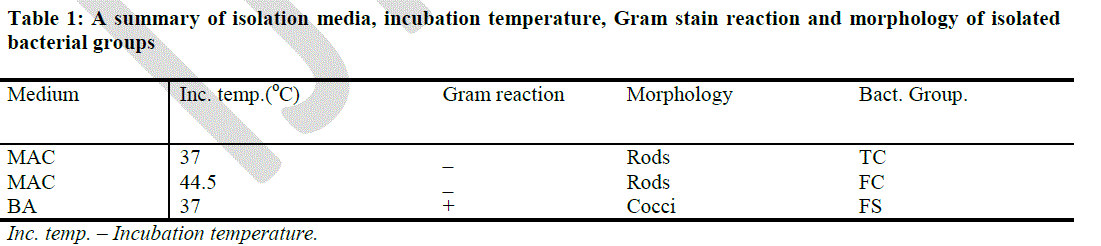
| A summary of the isolation media, incubation temperature, Gram stain reaction and morphology of isolated bacteria as seen under the microscope is shown in Table 1. The initial isolation of the three major bacterial groups under study was done on BA and MAC agars. Gram positive bacteria appeared bluish under the microscope while Gram negative bacteria appeared red under the microscope. Gram negative rods growing on MAC agar (37oC) after 48 hours were identified generally as the TC while Gram negative rods growing on MAC agar (44.5oC) after 48 hours were identified as the FC Gram positive culture isolates on BA agar (37oC) after 48 hours were identified as the FS. The colonies of TC and FC that were isolated on MAC agar plates were pinkish and reddish with smooth edges while the colonies of FS that were isolated on BA agar appeared dark or black. |
 |
| Results on catalase test were used to determine whether BA isolates were FS or non-FS bacteria. Isolates that produced bubbles upon addition of H2O2 were identified as catalase positive and were indicative of non-FS cocci. Isolates that did not produce bubbles upon addition of H2O2 were identified as catalase negative organisms and since the Gram stain reactions and microscopy had been done prior to the catalase test, isolates were identified as FS or non-FS bacteria. |
 |
| The results on the tests performed during isolation and identification of the FS bacteria from filtered and unfiltered effluents are shown in Table 2. The FS bacteria were found to be catalase negative, Gram positive cocci capable of aesculin hydrolysis, bile tolerance and citrate hydrolysis, the last three qualities being observed in their ability to grow on the BA agar. |
V. DETERMINATION OF NUMBERS OF TOTAL COLIFORMS, FAECAL COLIFORMS, FAECAL STREPTOCOCCI AND TOTAL BACTERIAL COUNTS IN EFFLUENTS. |
| Enumeration of bacteria was done in order to assess the effect of sand bed depth in reduction of the numbers of these bacterial groups in the filtered effluents. Results of bacterial counts in the filtered effluents at various sand bed depths showed that sand bed depth had no significant effect on reduction of bacterial counts (Fig. 2). Total coliform counts were found to be 6800 CFUs/ml in unfiltered effluents and reduced significantly (p < 0.05) to below 1000 CFUs/ml after filtration in all the three levels of experimental depths (Fig. 2 A). However, there was no significant difference in reduction of TC counts at the three levels of experimental depths and the average reduction of TC was 96.29 % (Fig. 2 A; Table 3). Faecal coliform counts were found to be 1900 CFUs/ml in unfiltered effluents and reduced significantly (p < 0.05) to below 500 CFUs/ml after filtration in all three experimental depths. There was no significant difference observed in FC reduction at depths 0.5 m (93%) and 0.7 m (94%). Interestingly, at 1.0 m depth, Percent removal was found to be lower at approximately 83% (Fig. 2 B; Table 3). |
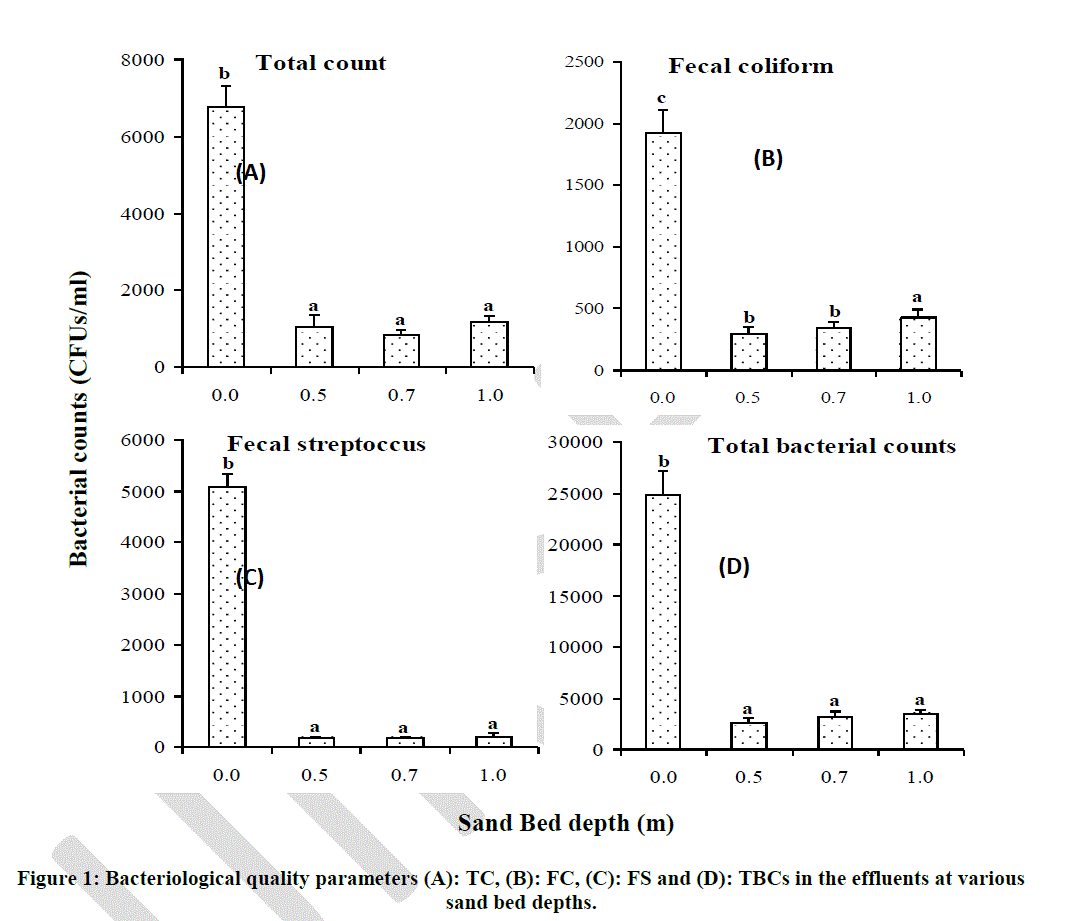 |
| Bars with the same letter superscript in the same graph are not significantly different at p < 0.05 Faecal coliform removal at 1.0 m sand bed depth was found to be 82.81% which was significantly different from removal at 0.5 m and 0.7 m depths (Table 10). Faecal streptococci had a similar trend to that of TC counts at the various sand bed depths, with an average removal of 98.12% (Fig. 2 C; Table 3). |
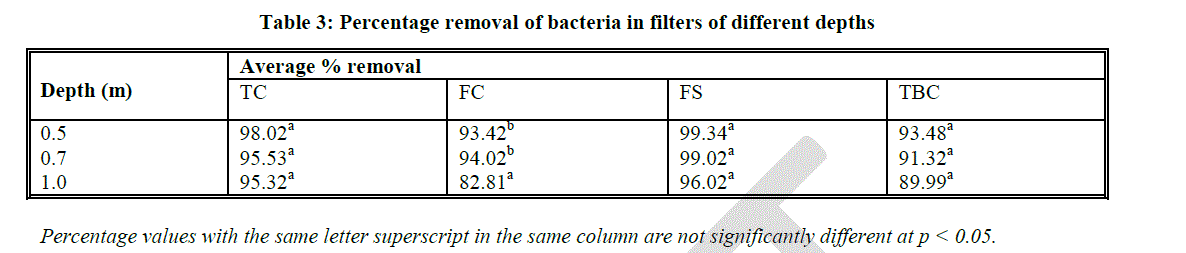 |
| Total bacterial counts were found to be 25000 CFUs/ml in unfiltered effluents and reduced significantly (p < 0.05) to below 5000 CFUs/ml after filtration in all the three experimental depths. However, just like for TC and FS counts, TBC reduction was not significantly different at the three levels of depth (Fig. 2 D). |
VI. PHYSICOCHEMICAL ANALYSES |
| Sand bed depth had a significant effect on pH, DO, BOD5, NO2 -, NO3 -, PO4 -3, conductivity and TSS (Figure 2). Although pH of the effluents was lowered significantly with increasing sand bed depth, there was no significant difference in filtered effluent pH at depths 0.5 m and 0.7 m. However, the pH at these two depths (0.5 m and 0.7 m) differed significantly from that obtained at depth 1.0 m which showed the lowest pH of 7.35 (Fig. 2 A). The DO of effluents increased significantly (p < 0.05) in filtered effluents as compared with the unfiltered effluents. Although, there were no significant differences in DO increment at the three levels of depth, increase in DO was found to be highest (7.60 mg/l) at depth 0.7 m and lowest (6.50 mg/l) at depth 1.0 m (Fig. 2 B). The BOD5 decreased significantly (p < 0.05) with an increase in sand bed depth. The BOD5was found to be highest in unfiltered effluents but lower at all the three levels of experimental depths. Significant differences in BOD5 were also observed among the three experimental depths (0.5 m, 0.7 m and 1.0 m) with the highest reduction being observed at 1.0 m depth and lowest reduction at 0.5 m depth (Fig 2 C). |
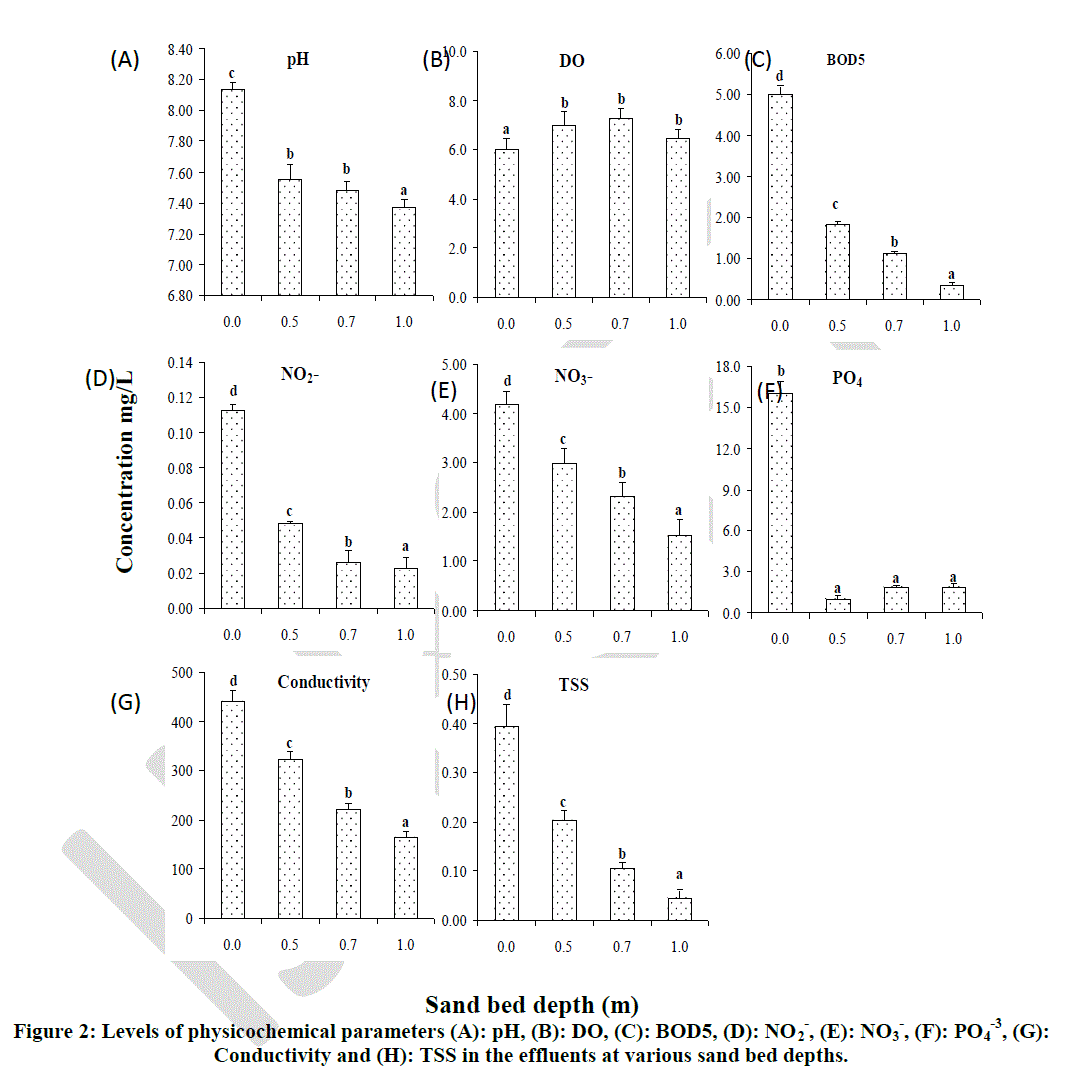 |
| Bars with the same letter superscript on the same graph are not significantly different at p < 0.05 |
VII. DISCUSSION |
| In slow SSF, the vertical height of the sand bed that the water has to pass through is important in terms of filtration efficiency. The reasons for this are the existence of biological activity in a sand filter which is known to occur at depths of up to 0.5 m within a sand bed and the available surface area for mechanical filtration [16]. |
| The removal rates of bacteriological and physicochemical parameters observed during this particular study are consistent with those of other studies using intermittent sand filters in the treatment of wastewater [18]. Elliot et al., [17], using laboratory scale model showed that longer depths of SSFs can be applied effectively as tertiary treatment for secondary effluents. The author reported that most of the removal of suspended solids and BOD5 occurred at the lower sand layer. The results in this study also showed that BOD5, NO2 -, NO3 -, Conductivity and TSS were significantly higher (p < 0.05) in unfiltered effluents but decreased significantly with increasing depth and the values for these parameters were found to be lowest at depths of 1.0 m. This showed that sand bed depth affected the reduction of the physicochemical parameters under study. These findings also agree with those of Bellamy et al.,[19] who reported an average percent removal of pH at 97%, BOD5 at 87%, NO3 - at 56%, and conductivity at 62% using sand bed depth of 1 m. |
| The findings in this study showed that PO4 -3 reduction was not significantly different at the different levels of sand depth. The low reduction capacity of PO4 -3 suggests that biomass communities within the sand bed removed negligible phosphorus from the wastewater [20]. Removal of phosphorus within sand beds is predominantly achieved through adsorption onto substratum and precipitation/fixation reactions [20]. Consequently, the surface area for chemistry can substantially govern the rate of phosphorus removal [20, 21, 22]. Although no chemical analysis of the substrate composition was undertaken, it can be assumed that it would be composed primarily of somewhat inert polymer chains of carbon and hydrogen that would have limited potential to bind phosphorus. Burgoon et al., [23] reported a maximum TP load removal rate of 44% in sand beds utilizing plastic tricking filter medium to treat primary treated wastewater, noting substrate adsorption and sedimentation as the major mechanism for the removal of TP. |
| The present study also sought to find out the role of sand bed depth on removal of TC, FC, FS and TBC and established that removal efficiency of these bacterial groups was not entirely pegged on sand bed depth. Results from this study showed that although maximum removal of the coliform organisms (98%) occurred at a depth of 0.5 m, there was no significant difference in bacterial removal in effluents at the different levels of sand bed depth. These results are consistent with those of Bellamy et al., [19] who reported a 97% removal of coliform bacteria at a sand depth of 0.9 m. |
| In a follow up study, Bellamy et al., [19] also reported 88% - 91% removal of standard plate counts at a depth of 1.0 m. These findings concur with those of our study that showed 90% removal of TBCs at the same sand bed depth. In addition, the present study established that percent removal of FC, TC and TBCs were not significantly different at 0.5 m, 0.7 m and 1.0 m depths. This confirms that bacterial removal in SSFs is not sensitive to sand bed depth. Likewise, Bellamy et al., [19] also found that removal of standard plate counts still ranged from 88% to 91 % when the sand depth was increased from 0.35 to 1.0 m suggesting that sand depth could be reduced to 0.48 m and still produce satisfactory bacteriological removal efficiency. |
| It can be hypothesized that sand bed depth is less significant in removal of bacteria. This is because most of the biomass and biological treatment occurs in the upper portion of the sand bed and the increasing depth would therefore have little effect on the filtered effluent quality in terms of bacterial removal [20]. For instance, Williams [24] found that all bacterial reduction occurs in the top 0.2 m of the filter bed. Research by ASCE, [25] also confirmed that majority of biological processes occur in the top 0.4 m of the sand bed. Bellamy et al., [19] and Muhammad et al., [26] reported that bacteriological treatment was not highly sensitive to sand bed depth. However, while this is generally true, bacteriological treatment efficiency becomes sensitive to bed depth with larger sand sizes. This is because the total surface area within the filter is reduced in a sand bed with larger grains and higher flow rates also occur, potentially increasing percolation rates [27]. |
| Wheelis [14] revealed interesting trends in bacterial removal in SSFs. This author reported 98%, 74% and 85% FS, FC and total plate count removals respectively at sand bed depths of 0.5 m. These results showed that FS removals were highest in SSFs followed by total plate count removals and the least removal was observed by FC. These findings coincide with those in the present study that revealed approximately 99%, 93%, 93% reductions of FS, FC and TBC respectively at the same sand bed depth. All these results portray satisfactory bacterial removals at a very small sand bed depth of 0.5 m showing that shallow bed depth probably allows more oxygen to diffuse to the microbes and the biologically active zone can grow deeper within the sand bed. |
VIII. CONCLUSION |
| From this study, it can be concluded that sand bed depth had a significant effect on removal of physicochemical parameters but did not influence the removal of bacteria in filtered effluents significantly. Physicochemical parameters such as pH, BOD5, NO3 -, NO2 -, conductivity and TSS are decreased significantly when sand bed depth is increased. However, reduction of PO4 -3 in filtered effluents is not significantly affected by sand bed depth. |
IX. ACKNOWLEDGEMENT |
| This work was supported by the National Council for Science and Technology (NCST), Kenya now National Commission for Science and Technology (NACOSTI) under Grant number NCST/5/003/POST-DOC/1ST CALL/015. |
References |
|