E- ISSN: 2320 - 3528
P- ISSN: 2347 - 2286
E- ISSN: 2320 - 3528
P- ISSN: 2347 - 2286
Dipto Sinha*
Department of Biotechnology, Grenoble Alpes University, Grenoble, France
Received: 07-Nov-2022, Manuscript No. JMB-22-79240; Editor assigned: 10-Nov-2022, PreQC No. JMB-22-79240(PQ);Reviewed: 24-Nov-2022, QC No. JMB-22-79240; Revised: 01-Dec-2022, Manuscript No. JMB-22-79240 (R); Published: 08-Dec-2022, DOI: 10.4172/2320-3528.11.7.001.
Visit for more related articles at Research & Reviews: Journal of Microbiology and Biotechnology
Cellular homeostasis is quintessential of regulated protein turnover machinery within a cell. Any form of interference with this highly orchestrated phenomenon of protein production in the Endoplasmic Reticulum (ER) can induce cellular stress. The continued blockade of the ER with misfolded proteins can lead to structural abnormalities of the ER and the mitochondria, following which a cascade of cellular reactions is triggered, marked by the increase in the levels of cytosolic Ca++ and dissipation of mitochondrial membrane potential. BAX promotes the loss of membrane potential of the mitochondria causing the release of Cytochrome-c and to counter it, BAX-inhibitor (BI-1) plays a crucial role in protecting the cell from undergoing Programmed Cell Death (PCD). A homolog of BI-1 from Plasmodium Spp. has been identified in the Toxoplasma genome (TgBI-1), which we intend to characterize by inducing stress on the parasite cell and then by reversibly modulating the levels of the concerned protein (putative BI-1) by using a tunable system like transactivator (Tet-off) to observe the repercussions in the cellular behaviour.
Toxoplasma gondii; BAX Inhibitor-1; Programmed Cell Death; Fluorescent microscope; Cytochrome-c
The obligate unicellular parasite Toxoplasma gondii can be quite ghastly as it has a very broad range of hosts [1]. During pregnancy it can cause foetal death or can inflict severe congenital defects in the foetus [2]. In immunocompromised individuals, the protozoan parasite can play havoc by causing Toxoplasmic encephalitis [3]. However, the successful parasite has to maintain a constant homeostasis to ensure its survival and to proliferate and propagate from one host to the other. The homeostasis within the cell is primarily dependant on the protein machinery and protein-folding. As the parasite gets to experience extremes of host cell environment change, undergoes a high rate of replication and the at the same time the parasite has to keep a repertoire of proteins, ready that are to be exported into the host cell, a constant burden on the ER is maintained. Nevertheless, a bulk of misfolded proteins also gets synthesized and the 26S proteasome machinery, in eukaryotic cell, clears it away. This heavy duty often leads to the inhibition of the proteasome and as a result of which misfolded proteins accumulate in the ER, causing ER stress. This, on one hand triggers ER stress response to salvage the parasite cell under stress or on the other hand, it can set the pathway for apoptosis on, usually when the content of misfolded proteins remain adherent in the ER for a long period of time [4,5]. Apoptosis (Programmed Cell Death-PCD) is marked by the release of Cytochrome-c through the pores of outer membrane of the mitochondrion of the cell. This entire episode is tightly regulated by the some member proteins that either supports it, like BAX (pore forming proteins) or hinders PCD, like Bcl-2. Recently, the screening of yeast proteome led to the finding of another antagonist of BAX, named BAX-inibitor-1 (BI-1) [6]. BI-1 is localized primarily in the ER and in the nuclear envelope as well, interestingly, it doesn’t interact with BAX but it can interact with Bcl-2 [7].
The presence of a putative BAX inhibitor candidate (Pf BI-1) (Gene ID–PF14_0571) in Plasmodium Spp. was reported to be synthesized during the asexual stages of the parasite development and proliferation which is when the cell is under tremendous pressure to maintain the cellular homeostasis [8]. A homologue of this Plasmodium BI-1 is found to exist in the Toxoplasma genome (Tg BI-1). We plan to unravel the role of this putative BI-1 (Tg BI-1) under the stress and normal conditions of the parasite cell by reversible expression of the gene (BI-1) using ATc regulated transactivator [9].
The BLASTp search of putative PfBI-1 against the Toxoplasma genome database (www.toxodb.org) led us to the identification of the putative BI-1 (Toxo Gene ID–TGDOM2_255900).
Toxoplasma gondii tachyzoites were grown by serial passage in Human Foreskin Fibroblast (HFF) cells as previously described [11]. The positive selection of the transfected parasites was made using chloramphenicol (20 uM concentration) as the vector carries a CAT (Chloramphenicol Acetyltransferase) cassette in its backbone followed by a clonal selection of the chloramphenicol resistant parasites (Supplementary Figure 1B) [12].
The amino acid residues of BI-1 from species Saccharomyces spp., Toxoplasma spp., Arabidopcis spp., Mus musculus and Homo sapiens were submitted to multiple sequence alignment site (https://www.ebi.ac.uk/Tools/msa/clustalo/) Figure 1A.
To determine the topology of the TgBI-1 protein, amino acid residue stretch was submitted to PepCal-C website (https://pepcalc.com/) Figure 1B. The residues projecting downwards mark the hydrophobic ones and these clearly denote and form the part of the transmembrane functional domains. The C-terminus however has charged amino acid residues like Lysine (denoted with blue bars) making the C-terminus hydrophilic and the possibility for the TgBI-1 C-terminus to function like a pH sensor.
The precise positioning of N-terminus of TgBI-1 was deduced by TMpred (www.ch.embnet.org/software/TMPRED form.html) Figure 1C and simultaneously TMHMM (www.cbs.dtu.dk/services/TMHMM) was used to do the same, Figure 1D.
Figure 1D: TMHMM prediction of TgBI-1 topology: TMHMM predicts the N-terminus of TgBI-1 to be jutting out of the ER lumen into the cytosol. As elucidated the in the TMHMM model, the transmembrane stretches span across are constituted of prominently hydrophobic amino acid residues and the N-terminus is mildly hydrophobic.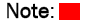

Localization of TgBI-1 was determined by in silico tool BUSCA (http://busca.biocomp.unibo.it/) Figure 1E.
Figure 1E: Prediction of intracellular localization of TgBI-1: BUSCA online tool predicts TgBI-1 to be targeted to the ER of the cell. TgBI-1 carries a consensus motif found exclusively in the ER resident proteins. Transmembrane (TM) helices predicted to be present and they mark the conserved and hydrophobic stretches in the molecule.
Free Toxoplasma gondii tachyzoites and intra-vacuolar parasites, formed in Human Foreskin Fibroblasts (HFF) cells were stained with DAPI (4',6-Diamidino-2-phenylindole dihydrochloride, 2-(4-Amidinophenyl)-6-indolecarbamidine dihydrochloride, DAPI dihydrochloride (to stain the nuclei)) were visualised using 100X objective on an Axioplan II microscope (Zeiss). Images were acquired using a Zeiss camera and the Axio Vision LE Rel. 4.5 software (release 4.7.1) Figure 1F and Figure 1G.
Figure 1F: Live cell microscopy of the free tachyzoites: Freshly egressed tachyzoites of Toxoplasma gondii were observed under fluorescent microscope. The DAPI stained nuclei of the parasite are seen encompassed by ER resident mCherry-myc-TgBI-1 protein, displaying red fluorescence. Inset (zoom) of the same further highlights the finding (DIC–Differential Interference Contrast).
Confluent HFFs were grown on glass coverslips and infected overnight with parasites. Infected cells were fixed in 5% formaldehyde, permeabilized with 0.002% saponin and blocked in 5% goat serum-5% FBS (in Phosphate Buffered Saline (PBS)). Cells were then incubated with primary antibodies for 1 hr in 1% FBS, 0.002% saponin. Primary antibodies used in this study included anti-GFP monoclonal antibodies (1:500) raised in mouse and anti-mCherry (1:500) antibodies raised in rabbit. Infected cells were washed and incubated for 1 hr with the following secondary antibodies: Goat anti-mouse IgG (H+L)-Alexa 488 (1:500, Jackson) and goat anti-rabbit IgG (H+L) Alexa 594 (1:500, Jackson). Coverslips were then incubated with 5 μg/ml Hoechst 33342 for 10 min to stain nuclei, mounted in Mowiol and observed using a 100X objective on an Axioplan II microscope (Zeiss). Images were acquired using a Zeiss camera and the Axio Vision LE Rel.4.5 software (release 4.7.1) Figure 1H.
Figure 1H: Immunofluorescence assay to confirm TgBI-1 in the ER: Intravacuolar parasites (mCherry-myc-TgBI-1 re-transfected with Der1-ER1-GFP) were stained with antibodies directed against mCherry (to recognize mCherry tagged TgBI-1) and GFP (to identify Der1-ER1 tagged with GFP). The complete overlap of the staining clearly establishes the localization TgBI-1 in the ER of the parasite cell.
The inset of the cropped image shows the individual staining of the two antibodies. (i) mCherry Antibodies: Distinctly mark the mCherry-mycTgBI-1. (ii) GFP antibodies: Der1 ER1 gets marked with the antibodies against GFP. (iii) Overlap of the two completely mask each other.
Topology, localization, functional domains and putative biochemical activity of the TgBI-1
Initially identified as a suppressor of BAX mediated cell death in yeast or mammalian cells, BAX Inhibitor-1 (BI-1) carries several evolutionarily conserved transmembrane domains and is predominantly localized in the Endoplasmic Reticulum [6,7,13-15].
Sequence analysis and topology of TgBI-1
BI-1 proteins are relatively small in molecular weight (25 kDa-41 kDa). As shown in Figure 1A-MSA, the domains of the proteins across different species remain well conserved as they show high degree of similarity in the entire sequences spread across different species.
The precise topology of the TgBI-1 remains unsolved, however bioinformatic analysis (https://pepcalc.com/) suggests the presence of six to seven transmembrane domains and the last transmembrane domain is less hydrophobic than the rest (Figure 1B). The C-terminus of TgBI-1, rich in lysine and glutamic acid residues (DKERRKRQRDEE), bears a passing resemblance with the consensus motif defined for type-I transmembrane ER resident proteins marked with the presence of lysine residues in the C-terminal end of the cytoplasmic domain [16].
The N-terminus positioning of TgBI-1 is still not very clearly determined. A model of TgBI-1 supported by TMpred (www.ch.embnet.org/software/TMPRED form.html), elucidates TgBI-1 N-terminus is buried in the ER and the semi-hydrophobic seventh transmembrane domain swoops into the membrane forming a loop domain which could be a reminiscent of voltage gated sodium channel SCN5A, suggesting the role of TgBI-1 as a channel protein (Figure 1C–TMpred) [17]. The extreme C-terminus end of TgBI-1 is rich in lysine residues (DKERRKRQRDEE) and acts as pH sensor in the BI-1 protein molecule and helps in tetramerization and enables the C-terminus to function as a Ca++ ion channel [18]. However, another model of the same to predicts the N-terminus as cytosolic, TMHMM (www.cbs.dtu.dk/services/TMHMM). N-terminus of BI-1 in yeast is reported to interact with an anti-apoptotic protein, NleH from Escherichia coli pointing at the significance of cytosolic localization of the N-terminus of the TgBI-1 (Figure 1D–TMHMM) this finding wasn’t observed in the mammalian cells though [19]. Here, we tagged the N-terminus of TgBI-1 with myc-mCherry as the protein is devoid of signal peptide sequence.
TgBI-1 is localized in the ER of the cell
Immunocytochemistry studies haven’t shed much light on localization of BI-1. Owing to the extreme hydrophobicity of the protein molecule, generation of reliable antibodies has not been very successful [6]. Nevertheless, in silico tool (http://busca.biocomp.unibo.it/) predicts TgBI-1 being targeted to the endomembrane system of the cell (Figure 1E, BUSCA) [20]. A Tetracycline-inducible Transactivator (TATi) system was employed in our study to transcriptionally regulate and study the localization of TgBI-1 upon being expressed [21]. The transgenic parasites (Figure 1I) were studied for the localization of mChery-myc-TgBI-1 fusion protein by fluorescent microscopy.
Figure 1I: Tagging the endogenous loci of TgBI-1-Schematic representation: Using a double cross-over approach the endogenous loci of the gene is replaced with a stretch from 5’UTR (marked with brown) and a fragment of the coding region (highlighted with blue) sandwiched in between the selection marker cassette (Chloramphenicol resistance cassette), Tet response element, visual marker–mCherry and myc tag. The protein bears TM domains as marked using brown colour.
Live cell imaging of the intracellular parasites and free tachyzoites displays the fluorescence of the tagged protein contained in the ER, encompassing the DAPI stained nuclei of the cells (Figure 1F and Figure 1G). The transcriptional downregulation of the gene product wasn’t observed (data not shown) possibly due to the residual expression of the gene [22]. To further confirm the localization of the tagged protein in the ER of the parasite cell, the transgenic TgBI-1-myc-mCherry cell-line was re-transfected with a vector bearing Der1-ER1, an ER resident protein, tagged with efGFP. In immunofluorescent assay, the staining of the two tagged proteins using anti-myc and anti-GFP antibodies overlapped confirming the localization of TgBI-1 in the ER of the parasite cell (Figure 1H).
Phylogenetic analysis of BI-1
Perhaps the most noteworthy fact about BI-1 is that it has remained evolutionarily well conserved. The negative regulator of apoptosis, BI-1 carries a signature conserved motif, UPF0005 that is ubiquitous across all kingdoms [23]. Functionally, UPF0005 is still uncharacterized and is present in cytoprotective proteins like BI-1, h-GAAP, Lifeguard (LFG). Owing to the hydrophobic nature of UPF0005, it is predicted to display six to seven transmembrane spanners and bioinformatic assessments indicate its localization in the endomembrane system of the cell and to be involved in inhibiting BAX induced cell death. Further sequence conservation analysis of UPF0005 support that BI-1, LFG and some closely related proteins constitute a family of six proteins with anti-apoptotic role, called the BI-1 family or Transmembrane BAX Inhibitor Motif containing (TMBIM) [6].
TMBIM1/RECS1: Initially identified as a gene playing proactive role in response to centrifugal force and shear stress, hence the name RECS, is a 35 kDa protein and is ubiquitously expressed in all cells expect for thymus, spleen and testis [24,25]. It is reported to be localized in the membranous compartments like plasma membrane, endosomes, lysosomes and later was established to be found in golgi apparatus as well [25,26]. RECS1, also known as PP1201, LFG3, MST100, MSTP100 protects the cell from Fas mediated programmed cell death by reducing the
distribution of Fas on the cell surface and at molecular level TMBIM1/RECS1 forms a complex with receptor Fas/CD95/Apo1 at golgi apparatus [26].
TMBIM2/LFG2: In a genetic screening procedure to identify genes that can block cell death induced by Fas ligand exclusively with little or no role to play by TNFa, TMBIM2 also known as NMP35, FAIM2, LFG2, NGP35, KIAA0950 was seen to play a pivotal role [27]. TMBIM2/NMP35 is a 35 kDa protein which displays a neuronal expression pattern, predominantly in the dendritic processes and in a subset of synapses at the postsynaptic membrane and more prominently in the ER, golgi apparatus and lipid raft micro domains of the plasma membrane [27-29].
TMBIM3 also known as LFG1, HNRGW, NMDARA1, MGC99687, GRINA, GBP is approximately a 41 kDa protein with seven transmembrane helices, has been reported in the marginal zone of the cerebral cortex of developing mouse brain. Information pertaining to this protein isn’t very abundant, however, it was initially identified as a candidate protein displaying activity against antibody raised against glutamate-binding protein, a part of an NMDA receptor-associated complex [30,31].
TMBIM4 was identified as a human golgi anti-apoptotic protein with a highly conserved homologue in vaccinia virus, hence the name GAAP [32]. Also referred to as S1R, LFG4, ZPRO, CGI-119, TMBIM4/GAAP is a 27 kDa protein with seven transmembrane domains and localizes in the ER and golgi apparatus membranes. It is known to lower the Ca++ levels in the intracellular stores which reduces the efficiency of IP3 which comes into play when histamine induced Ca++ release is triggered, thereby making the cell resilient to PCD [33]. Overexpression of GAAP inhibits apoptosis induced by Tumour Necrosis Factor (TNF) α and FasL [32].
TMBIM5, a 37 kDa protein predicted to bear eight transmembrane domains is the only member of this family of proteins that is localized in the inner membrane of mitochondria [34,35]. It earned the name GHITM as its mRNA was noticed to be down regulated in transgenic mouse expressing a growth hormone antagonist [36]. Functional analysis studies of this gene reveals that knocking it out leaves the mitochondrial cristae disorganized and mitochondria is rendered fragmented. TMBIM5 prevents the release of Cytochrome-c from the mitochondria upon treatment of Actinomycin D, which induces mitochondrial fragmentation [35].
Function of BI-1
Interaction with BCL-2 family of proteins and role in PCD: Studies carried out on BI-1 clearly establish that it protects the cells from undergoing PCD. The initial work done on yeast cells show that the BI-1 protected the cells when human pro-apoptotic protein BAX was over-expressed. BI-1 served the same function when apoptosis inducer BAX was over-expressed human HEK293 cells as well [6]. The function of BI-1 is extremely conserved as elucidated when BI-1 from plant origin (Oryza sativa and Arabidopsis thaliana) could avert the PCD induced by BAX and hydrogen peroxide in yeast cells [37]. BI-1 interacts with BCL-2 family of proteins, BCL-2 and BCL-xL which was established by co-immunoprecipitation upon cross-linking the over expressed BCL-2 and BI-1 [6]. BCL-2, a ER membrane and nuclear envelope resident protein and BCL-xL, a mitochondrial membrane protein influence the Ca++ concentration in the ER [38]. BCL-xL-1 alone can’t bring about the depletion of ER (Ca++) when BI-1 is knocked down using hairpin RNA against BI-1. It instead, leads to dramatic increase in the ER (Ca++) thereby re-establishing the pivotal role played by BI-1 in maintaining the ER homeostasis [39].
BI-1 mediates the level of Cytochrome P450 2E1: Lipid peroxidation of the ER membrane brought in by Reactive Oxygen Species (ROS) is often triggered by ER stress. The over-expression of BI-1 considerably brings down the level of oxidation damage and protects the ER membrane from the harm. BI-1 regulates the Cytochrome P450 2E1 levels and significantly reduces the ROS production level by Cytochrome P450 2E1 generated in the liposomes [40]. BI-1 is not known to interact directly with Cytochrome P450 2E1, however, findings suggest that BI-1 does physically interact with another member of the microsomal monooxygenase system, NADPH-P450 reductase. Studies indicate that BI-1 induces disruption of interaction between NADPH-P450 reductase and Cytochrome P450 2E1, halting the transfer of electrons between the two proteins and eventually bringing down the ROS levels. The inhibitory effect of BI-1 against damage by ROS is ceased with the addition of antibodies raised against the C- terminus of BI-1 or when BI-1 with abrogated C-terminus is expressed, hinting at the key role played by the highly conserved C-terminus amino acid residues [40].
The C-terminus of BI-1 functions as Ca++ channel pore that mediates Ca++ leak from ER into the cytosol. This Ca++ flux property of the BI-1 C-terminus is unique and is different from the other classically defined Ca++ channels. Experiments done using C-terminal peptide of BI-1 on artificial lipid bilayer membrane showed that it forms a loop like structure, whereas adding inhibitors against intracellular Ca2++ release/leak channels like IP3Rs, RyRs doesnot hamper the property of the BI-1 to function as a Ca++ channel. Furthermore when aspartic acid residue at 213 is replaced with arginine in full length BI-1 in HeLa cells, BI-1 fails to initiate the efflux of Ca++ from ER into the cytosol [15]. The TgBI-1 C-terminus amino acid residues (DKERRKRQRDEE) display a strong degree of homology with the BI-1 found and characterized in other species. This strongly indicates that the TgBI-1 could also be implicated in the maintenance of Ca++ balance in the ER and functions as Ca++ anti-porter as well [41-46].
Previously BI-1 was identified as Testis Enhanced Gene Transcript (TEGT) in mammals owing to its abundance in testis. The negative regulator of cell death, BI-1 (TMBIM) family of proteins remains evolutionarily highly conserved and have been reported to be present in species where BCL-2 family of proteins have not been discovered yet including some plants, bacteria and even viruses. Some recent studies establish the localization of BCL-2 proteins in previously unreported organelles where they are involved in specific novel functions. With the significant progress made in understanding the functioning of TMBIM family of proteins we now know that BI-1 plays a key role in calcium homeostasis, the UPR, lysosomal function, autophagy and mitochondrial bioenergetics. This allows us to view BI-1 as a “stress sentinel” in the context of dealing with cell survival response by casting an antagonistic effect on apoptosis or PCD triggered by UPR even in case of apicomplexan parasites. However, the major hurdle that remains to be overcome is to create hierarchical map of the TMBIM family of proteins based on the biochemical level of functioning. With the lack of classical machinery for apoptosis in apicomplexan genome involving BCL-2 and the discovery of a member of TMBIM family of proteins namely, TgBI-1, it is quite likely that perspective of evolution of PCD, TMBIM group of proteins govern the PCD in a more fundamental and conserved manner in the apicomplexan and other lower species.
[Crossref][Google scholar][EBSCO]