Department of Statistics, University of Pune, Pune 411007, India
Received: 29/08/2015 Accepted: 30/12/2015 Published: 06/01/2016
Visit for more related articles at Research & Reviews: Journal of Statistics and Mathematical Sciences
Frailty models are used in survival analysis to model unobserved heterogeneity. To study such heterogeneity by the inclusion of a random term called the frailty is assumed to multiply hazards of all subjects in the shared frailty. We study compound negative binomial distribution as frailty distribution and two different baseline distributions namely Pareto and linear failure rate distribution in this paper. A simulation study is done to compare the true values of parameters with the estimated value. We develop the Bayesian estimation procedure using Markov Chain Monte Carlo (MCMC) technique to estimate the parameters of the proposed models. We try to fit the proposed models to a real life bivariate survival data set of McGrilchrist and Aisbett related to kidney infection. Also, we present a comparison study for the same data by using model selection criterion, and suggest a better model.
Bayesian estimation, Compound negative binomial frailty, Markov chain monte carlo (MCMC), Shared frailty.
Frailty is a random component designed to account for heterogeneity caused by unobserved individual-level factors that are otherwise neglected by the other predictors in the model. Vaupel et al. [1] suggested frailty models to account for the variations due to unobserved covariates. Several distributions like gamma, inverse Gaussian, positive stable distribution, power variance function, Weibull, compound poisson are used as frailty models for heterogeneity in the populations.
A class of random effect models which proved useful in survival analysis of related individuals is a class of frailty models which are based on the modifying the hazard function of individuals by introducing multiplicative effect on the baseline hazard function. Thus the frailty model is a random effect model for time to event data which is an extension of the Cox's proportional hazards model. Vaupel et al. [1], Keyfitz and Littman [2] showed that ignoring individual heterogeneity lead to incorrect conclusions.
Let T be a survival time with an absolutely continuous distribution. A non-negative random variable random variable Z is called ‘frailty’ if the conditional hazard functions given Z=z is given by
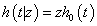 (1.1)
(1.1)
Where h0(t) is called the baseline hazard function. Then the conditional survival function is given by,
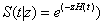 (1.2)
(1.2)
Where 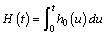 is the cumulative baseline hazard.
is the cumulative baseline hazard.
And the marginal survival function
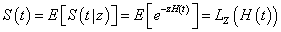 (1.3)
(1.3)
Where LZ (.) is the Laplace transform of the frailty distribution.
In this paper, we consider the shared frailty model with the compound negative binomial distribution as a frailty distribution and the Pareto and linear failure rate distribution as baseline distributions. The remainder of the paper is organized as follows. In Section 2, we give the properties of the general shared frailty models. In Section 3, we introduce the shared compound negative binomial frailty model. The baseline distributions are given in Section 4. In Section 5, we propose two different compound negative binomial frailty models. In Section 6, we discuss Bayesian method is used to estimate the parameters of the proposed models. We discuss the different model selection criterion by Bayesian approach in section 7. In Section 8 we present the simulation study. We present analysis of kidney infection data set and suggest a better model from these two proposed models in Section 9. Finally in Section 10, we discuss the conclusion of the study.
Shared frailty models explain correlations within groups (family, litter or clinic) or for recurrent events facing the same individual. i.e., the different events within each community share a common frailty, shared by each individual within the community and each unit belongs to precisely one category. The shared gamma frailty model was suggested by Clayton [3] for the analysis of the correlation between clustered survival times in genetic epidemiology.
Bivariate survival data arise when each subject understudy experiences two events. For e.g., failure times of paired organs like kidneys, eyes, ears or any other paired organs of an individual, In industrial applications, the breakdown times of dual generators in a power plant or failure times of two engines in a two-engine airplane, recurrences of a given disease. Both monozygotic and dizygotic twins share date of birth and common pre-birth environment are the illustrations of bivariate survival data with the shared frailty [4].
In order to build the shared frailty model for such kind of the data it is assumed that survival times are conditionally independent, for a given shared frailty. i.e., there is an association between survival times only due to frailty.
Let a bivariate random variable 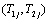 be the survival time of the ith ( i=1, 2) component of the jth individual (j=1,2,..,n). Given the unobserved Zj the hazard function for the
be the survival time of the ith ( i=1, 2) component of the jth individual (j=1,2,..,n). Given the unobserved Zj the hazard function for the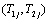 given by,
given by,
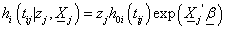 i=1,2 (2.1)
i=1,2 (2.1)
Where zj represents frailty acting as multiplicative effect at an individual level, 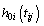 is the baseline hazard at time tij > 0 and
is the baseline hazard at time tij > 0 and 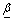 is the vector of regression coefficients with k components and
is the vector of regression coefficients with k components and 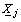 is the vector of observed covariates having k components.
is the vector of observed covariates having k components.
Integrating the hazard function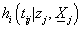 we get, the conditional cumulative hazard function for the jth individual at the ith component survival time tij > 0 for the given frailty
we get, the conditional cumulative hazard function for the jth individual at the ith component survival time tij > 0 for the given frailty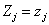
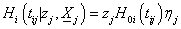 (2.2)
(2.2)
Where 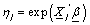 and
and 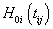 is the cumulative baseline hazard function at time tij > 0. The conditional survival function for the jth individual at the ith component survival time tij > 0 for the given frailty
is the cumulative baseline hazard function at time tij > 0. The conditional survival function for the jth individual at the ith component survival time tij > 0 for the given frailty 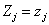 is,
is,
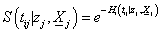
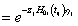
When 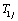 and
and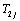 for j=1,2,..,n are independent, the bivariate conditional survival function of
for j=1,2,..,n are independent, the bivariate conditional survival function of 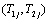 for the given frailty
for the given frailty 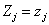 is the product to conditional survival function of
is the product to conditional survival function of 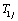 and
and 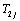 for the given frailty
for the given frailty 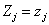 Thus, we have
Thus, we have
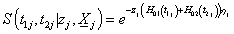
Where 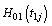 and
and 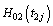 be the cumulative baseline hazard functions of the first and the second component at
be the cumulative baseline hazard functions of the first and the second component at 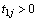 and
and 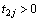 respectively.
respectively.
Integrate out the bivariate conditional survival function of 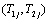 over the frailty variable Zj having the probability function, fz (Zj) for the jth individual in order to obtain the unconditional bivariate survival function at time Tij >0.
over the frailty variable Zj having the probability function, fz (Zj) for the jth individual in order to obtain the unconditional bivariate survival function at time Tij >0.
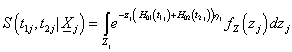
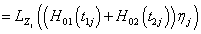
Where (.) LZ j is the Laplace transform of frailty the variable of zj for jth individual.
A compound distribution is a model for a random sum Z=X1+X2+…+XN where the number of terms N is uncertain. We assume that the variables Xi are independent and identically distributed and that each Xi is independent of N. The random sum Z can be interpreted the sum of all the measurements that are associated with certain events that occur during a fixed period of time. For example, compound distribution is the random variable of the aggregate claims generated by an insurance policy or a group of insurance policies during a fixed policy period. In this setting, N is the number of claims generated by the portfolio of insurance policies and X1 is the amount of the first claim and X2 is the amount of the second claim and so on. The random sum Z is said to have a compound Poisson distribution if N follows the Poisson distribution.
The Compound Poisson variable Z is defined as
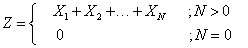
Where N is Poisson distributed with mean ρ while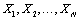 are independent gamma distributed with scale parameter v and shape parameterξ.
are independent gamma distributed with scale parameter v and shape parameterξ.
The distribution of Z can be partitioned into two parts, one a discrete part corresponding to the probability of zero susceptibility and second is due to continuous part on the positive real line.
The discrete part is given as
P(Z = 0) = exp(−ρ )
Which is decreases as ρ decreases.
The distribution of the continuous part is given by conditioning Z and with X having gamma distribution.
Aalen [5,6] introduced compound Poisson distribution as a mixing distribution in survival models. In many situations hazard rates or intensities are raising at the start, reaching a maximum value and then declining, that’s why the intensity has a unimodal shape with finite mode.
For e.g. death rates for cancer patients, divorce rates.
The reason to start decline in the population intensity is that the high risk individuals have already died or been divorced in case of above examples.
Also in is often seen that the total integral under the intensity or hazard rate is to be finite. It occurs due to the distribution is defective. It means that some individuals have zero susceptibility; they will survive forever.
In the case of above examples some patients survive their cancer, some people never marry, and some marriages are not prone to be dissolved. In such kind of data compound Poisson distribution plays an important role of mixing distribution.
Even though the compound Poisson distribution has many attractive properties, in non-susceptible or zero susceptibility type of data a convenient frailty distribution. Some other examples of zero susceptibility type of data if N > 0 then we can interpret Z as aggregate heterogeneity due to failures before we get first success or in general rth success can be given as, in case of marriage data, Z may represents heterogeneity due to difficulties in finding a marriageable partner before individual meet first suitable partner. In case of fertility, Z may be heterogeneity due to miss-carriages observed or unable to conceive a child with a couple before they have their first child or second child. Some mothers would like to deliver babies until she delivers a baby boy or two baby boys. Politicians go on contesting elections until they win once or twice and so on. Individuals go on changing the jobs until he/she gets a suitable job. In such situations negative binomial distribution or geometric distribution is a suitable choice of distribution for variate N.
So, we decide to consider compound negative binomial distribution as a frailty distribute-ion to model zero susceptibility type data in our study.
When the number of successes is equal to one, the compound negative binomial distribution reduces to compound geometric. There is a relation between compound negative binomial distribution and compound Poisson distribution. If Z1 and Z2 are respectively follows compound Poisson and compound negative binomial distribution with Z1=X1+X2+…+XN1 and Z2=X1+X2+… +XN2 where N1 is Poisson random variate with intensity λ and N2 is and negative binomial random variate with parameters, number of successes and probability
of success P then Z1 and Z2 are identically distributed if λ = −r log p .
The compound negative binomial distribution is defined as
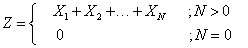
Where N is negative binomial variate with parameters r and p; r and p denotes respectively, the number of successes and the probability of success while X1,X2,…,XN are independent gamma distributed with scale parameter ν and shape parameter ξ.
The distribution of Z consists of two parts; a discrete part which corresponds to the probability of zero susceptibility, and a continuous part on the positive real line.
The discrete part is,
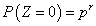
The distribution of the continuous part can be found by conditioning N and using the fact that the X's are gamma distributed. It can be written as
 (3.1)
(3.1)
is the density function of compound negative binomial distribution.
The expectation and variance of Z are
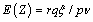 and
and 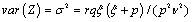
The Laplace transformation of Z is given by,
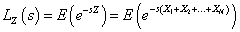
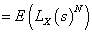
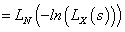
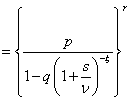
In order to solve the non-identifiability problem, we take E (z) =1 which leads to a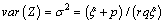 . Thus, we have the following form of Laplace transformation.
. Thus, we have the following form of Laplace transformation.
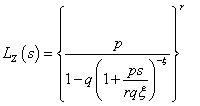
The unconditional bivariate survival function for the jth individual at the time t1j > 0 and t2j > 0 by replacing the above mentioned Laplace transformation in equation (2.5) we have,
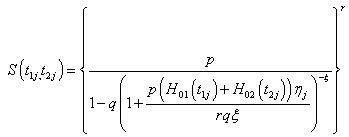
Where 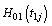 and
and 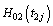 are the cumulative baseline hazard functions of lifetime random variables T1j and T2j respectively.
are the cumulative baseline hazard functions of lifetime random variables T1j and T2j respectively.
In this paper, we consider the two baseline distributions namely Pareto and linear failure rate distribution which yield two compound negative binomial frailty models.
The Pareto distribution is a skewed, heavy-tailed distribution that is sometimes used to model the distribution of incomes. This distribution is not limited to describing wealth or income, but to many situations in which an equilibrium is found in the distribution of the "small" to the "large".
The first baseline distribution is the Pareto distribution [7]. A continuous random variable T is said to follow the Pareto distribution with the scale parameter and the shape parameter α if its survival function is,
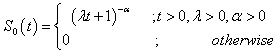 (4.10
(4.10
and the hazard function and the cumulative hazard function as
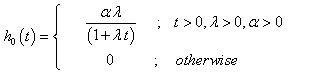 (4.2)
(4.2)
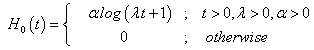 (4.3)
(4.3)
Observe that h0(t) decreases with t; λ> 0, α > 0. Hence this distribution belongs to the Decreasing Failure Rate class.
The exponential and Rayleigh are the two most commonly used distributions for analyzing lifetime data. These distributions have several desirable properties and nice physical interpretations. Unfortunately the exponential distribution only has constant failure rate and the Rayleigh distribution has increasing failure rate. The linear failure rate distribution generalizes both these distributions. We consider this is the second baseline distribution.
The linear failure rate distribution of a continuous random variable T with the parameters α >0 and λ >0, will be denoted by LFRD (α,λ) has the following survival function
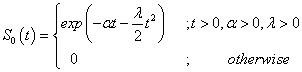 (4.4)
(4.4)
It is easily observed that the exponential distribution (ED (α)) and the Rayleigh distribution (RD (λ)) can be obtained from LFRD (a, b) by putting λ =0 and α =0 respectively. Moreover, the probability density function (PDF) of the LFRD (α,λ) can be decreasing or unimodal but the failure rate function is either constant or increasing only. See for example Bain [8], Sen and Bhattacharya [9], Lin et al. [10], Ghitany and Kotz [11] and the references cited therein.
The hazard function and the cumulative hazard function of linear failure rate distribution are respectively as stated below:
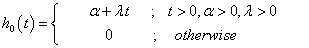 (4.5)
(4.5)
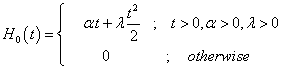 (4.6)
(4.6)
Here we present the two compound negative binomial frailty models say Model I and Model II by putting respectively the cumulative hazard function of the baseline distributions namely Pareto and linear failure rate distribution in the unconditional survival function of bivariate random variable 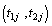 given in equations (4.3) and (4.6).
given in equations (4.3) and (4.6).
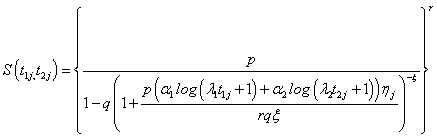
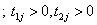 (5.1)
(5.1)
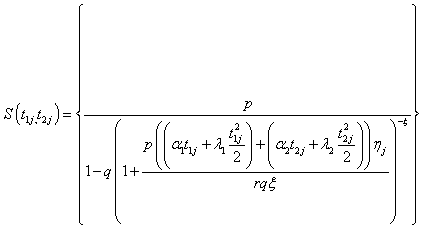
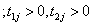 (5.2)
(5.2)
Here onwards, equations (5.1) and (5.2) as Model I and Model II which correspond to compound negative binomial frailty models with baseline Pareto and linear failure rate distributions respectively.
For the bivariate life time distribution, we use the bivariate censoring scheme given by Hanagal and Dabade [12].
Suppose that there are n independent pairs of components under study and the jth pair of the component have lifetimes 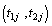 i.e. there are n individuals with a pair of components having lifetimes
i.e. there are n individuals with a pair of components having lifetimes 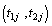 for j=1, 2,… , n. The life time times associated with jth individual is given by,
for j=1, 2,… , n. The life time times associated with jth individual is given by,
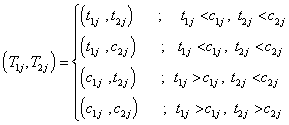
Where c1j and c2j be the observed censoring times for jth individual (j=1,2,…,n) with a pair of components respectively. We assume that the lifetimes and censoring times are independently distributed.
Now the likelihood of the sample of size n is given by,
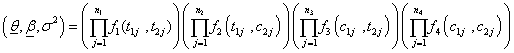 (6.1)
(6.1)
Where θ ,β ,σ2 are the vector of parameters of the baseline distributions, the vector of regression coefficients and the frailty parameter respectively. Let the counts n1,n2,n3 and n4 and be the number of individuals for which the first and the second components failure times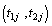 lie in the ranges;
lie in the ranges; 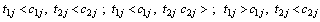 and
and 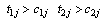 respectively and
respectively and
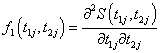
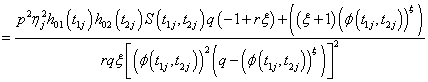
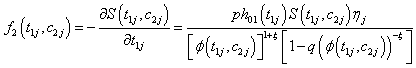
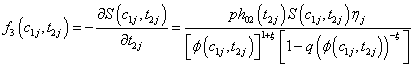
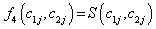 (6.2)
(6.2)
Where
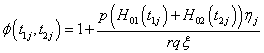
Thus, we get the two likelihood functions for the two proposed compound negative binomial frailty models namely Model I, Model II by substituting the corresponding hazard functions and cumulative hazard functions in the likelihood function given by equation (6.1) with 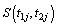 stated in equation (5.1) and (5.2).
stated in equation (5.1) and (5.2).
The likelihood function is obtained for the
Model I with
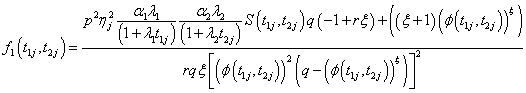
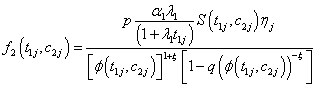
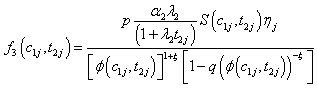
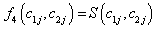
Where
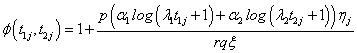 (6.3)
(6.3)
Model II with
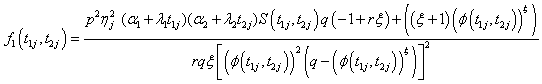
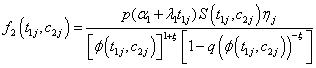
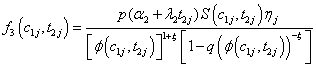
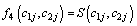
Where
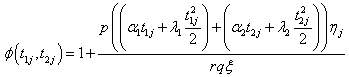 (6.4)
(6.4)
Each of the proposed models consists of eleven parameters, computing the maximum likelihood estimators (MLEs) involves solving a eleven dimensional optimization problem for these model. As the method of maximum likelihood fails to estimate the several parameters due to convergence problem in the iterative procedure, so we use the Bayesian approach. The traditional maximum likelihood approach to estimation is commonly used in survival analysis, but it can encounter difficulties with frailty models. Moreover, standard maximum likelihood based inference methods may not be suitable for small sample sizes or situations in which there is heavy censoring [13]. Thus, in our problem a Bayesian approach, which does not suffer from these difficulties, is a natural one, even though it is relatively computationally intensive.
Several authors have discussed Bayesian approach for the estimation of parameters of the frailty models. Some of them are, Ibrahim et al. [14] and Santos and Achcar [15]. Santos and Achcar [15] considered the parametric models with Weibull and the generalized gamma distribution as the baseline distributions and gamma and log-normal as frailty distributions. Ibrahim et al. [14] and references therein considered Weibull model and the piecewise exponential model with gamma frailty. Therefore we proposed Bayesian inferential approach in this study to estimate the parameters of the model, which is a popularly used method, because computation of the Bayesian analysis becomes feasible due to advances in computing technology.
To apply markov chain monte carlo (MCMC) methods, we assume that, conditional on observed covariates and on the entire set of parameters, observations are independent and prior distributions for all parameters are mutually independent. We used the Metropolis-Hastings algorithm within Gibbs sampler technique which is the most basic MCMC method used in Bayesian Inference. Convergence of Markov chain to a stationary distribution is observed by the trace plots, the coupling from the past plots, the Gelman-Rubin convergence statistic, and the Geweke test. The trace plots are used to check the behavior of the chain and the coupling from the past plots can be used to decide the burn-in period. The Gelman-Rubin convergence statistic values are approximately equal to one then sample can be considered to be come from the stationary distribution. The Geweke test examine the convergence of a Markov chain based on the sub parts of a chain at the end and at the beginning of the convergence period.
The large standardized difference between ergodic averages at the beginning and at the end of the convergence period indicates non convergence. The sample autocorrelation plots can be used to decide the autocorrelation lag.
In Bayesian paradigm the parameters of the model are viewed as random variables with some distribution known as prior distribution. It enables us to combine both the prior information and the data at hand to update the information of parameter. Thus, posterior density of a parameter is the distribution of a parameter updated by combining its prior distribution and the likelihood function. We assume that, conditional on explanatory variables and on the entire set of parameters, observations are independent and prior distributions for all parameters are mutually independent while applying MCMC methods.
Let 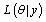 be the likelihood function and p(θ ) be the prior density of a parameter then posterior density function of a parameter π (θ | y) is given by,
be the likelihood function and p(θ ) be the prior density of a parameter then posterior density function of a parameter π (θ | y) is given by,
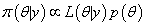 (6.5)
(6.5)
In our case the joint posterior density function of a parameter for given failure times 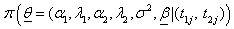
as
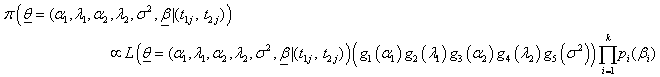 (6.6)
(6.6)
Where gi ( .) for i= 1, 2, ..,5 denotes the prior density function with known hyper parameters of corresponding argument for baseline parameters and frailty variance; pi (βi ) is the prior density function for regression coefficients βi for i =1, 2, ..,k.,. And the likelihood function L (.) is given by equation (6.1).
Algorithm consists in successively obtaining a sample from the conditional distribution of each of the parameter given all other parameters of the model. These distributions are known as full conditional distributions. Since the full conditional distributions are not easy to integrate out therefore full conditional distributions are obtained by considering that they are proportional to the joint distribution of the parameters of the model. The samples are then obtained from these full conditional distributions.
Bayesian model comparison is commonly performed by computing posterior model probabilities. In order to compare proposed models we use the akaike information criterion (AIC), the Bayesian information criterion (BIC), the deviance information criterion (DIC) and the Bayes factor. These are the most common methods of Bayesian model assessment.
Akaike [16] suggested that, given a class of competing models for a data set, one choose the model that minimizes
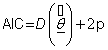 (7.1)
(7.1)
where p represents number of parameters of the model. 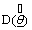 represents an estimate of the deviance evaluated at the posterior mean,
represents an estimate of the deviance evaluated at the posterior mean,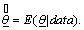 The deviance is given by, D(θ ) = −2 log L(θ) Where
The deviance is given by, D(θ ) = −2 log L(θ) Where 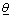 is a vector of unknown parameters of the model and L(θ ) is the likelihood function of the model.
is a vector of unknown parameters of the model and L(θ ) is the likelihood function of the model.
Bayesian information criterion (BIC) was suggested by Schwarz [17]. Shibata [18] and Katz [19] have shown that the AIC tends to overestimate the number of parameters needed, even asymptotically. The Schwarz criterion indicates that the model with the highest posterior probability is the one that minimizes
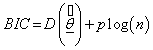 (7.2)
(7.2)
Where n is the number of observations, or equivalently, the sample size.
DIC, a generalization of AIC, is introduced by Spiegelhalter et al. [20] and is defined as;
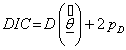 (7.3)
(7.3)
Where 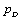 is the difference between the posterior mean of the deviance and the deviance of the posterior mean of parameters of interest, that is,
is the difference between the posterior mean of the deviance and the deviance of the posterior mean of parameters of interest, that is, 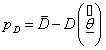 and
and 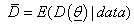
Models with smaller values of the AIC, BIC and DIC are preferred.
Kadane and Lazar [21] review model selection from Bayesian and frequent perspectives. The Bayes factor BJK for a model MJ against MK or given data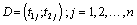 is
is
Where 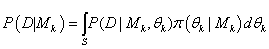
θk is the parameter vector under model MK and 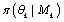 is prior density and s is the support of the parameter θk. Raftery [22], following Jeffreys [23], proposes the rules of thumb for interpreting twice the logarithm of the Bayes factor. For two models of substantive interest, MJ and MK twice the log of the Bayes factor is approximately equal to the difference in their BIC approximations.
is prior density and s is the support of the parameter θk. Raftery [22], following Jeffreys [23], proposes the rules of thumb for interpreting twice the logarithm of the Bayes factor. For two models of substantive interest, MJ and MK twice the log of the Bayes factor is approximately equal to the difference in their BIC approximations.
To compute Bayes factor we need to obtain 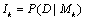 . By considering one of the approaches given in Kass and Raftery [24], we obtain the following MCMC estimate of IK which is given by,
. By considering one of the approaches given in Kass and Raftery [24], we obtain the following MCMC estimate of IK which is given by,
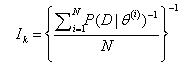 (7.5)
(7.5)
Which is harmonic mean of the likelihood values. Here N represents the posterior sample size and {θ(i),i = 1 , 2 ,..., N} is the sample from the prior distribution.
A simulation study is done to evaluate the performance of the Bayesian estimation procedure. For the simulation purpose we have considered only one covariate X1 and we assume that it follows normal distribution. The frailty variable Z is assumed to have compound negative binomial distribution. Life times (T1j,T2j) for the jth individual are conditionally independent for the given frailty Zj=zj. We considered that Tij (i=1,2;j=1,2,…,n) follows one of the baseline distributions, namely, Pareto (Model-I) or linear failure rate (Model-II) distribution respectively. As the Bayesian methods are time consuming, we generate only twenty, fourty and sixty pairs of lifetimes using inverse transform technique. Here we have generated different random samples of size n=20, 40 and 60 for lifetimes T1j and T2j. But here we are giving procedure for sample generation of only one sample size, say, n=20. Samples are generated using the following procedure:
1. Generate a random sample of size 20 from the compound negative binomial distribution with shape and the scale parameter ν as shared frailties (zj) for jth (j=1,2,…,20) individual.
To generate random observation from compound negative binomial distribution, we firstly generate a random observation N=n from negative binomial distribution with parameters
r=1, p=0.2.
Then we consider the following two cases:
(i) If N=0; take frailty Z=0.
(ii) If N>0; generate gamma variates say X1,X2,…,XN with the shape parameter ξ =0.2 and scale parameter 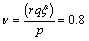 then frailty is taken as Z=X1+X2+…+XN.
then frailty is taken as Z=X1+X2+…+XN.
2. Generate 20 covariate values for X1 from the normal distribution.
3. Compute 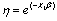 with the regression coefficient β=0.5.
with the regression coefficient β=0.5.
4. Generate 20 pairs of lifetimes (T1j, T2j) for the given frailty (zj) using the following generators,
for Model-I and Model-II respectively,
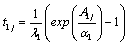 ,
, 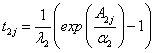 (8.1)
(8.1)
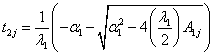 (8.2)
(8.2)
Where 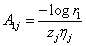 and
and 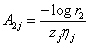 and rij i=1,2 are random sample from U (0,1).
and rij i=1,2 are random sample from U (0,1).
5. Generate the censoring times (c1j and c2j) from the exponential distribution.
6. Observe the ith survival time tij = min(tij, cij) and the censoring indicator δij for the jth individual (i= 1;2 and j=1,2,….,20), Where
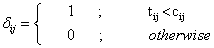
Thus we have data consists of 20 pairs of the survival times (T1j; t2j) and the censoring indicators δij .
We run two parallel chains for the proposed model with the different starting points using Metropolis-Hastings algorithm within Gibbs sampler based on normal transition kernels.
We iterate both the chains for 95,000 times. In our study we use non-informative prior for the frailty parameter σ2 and the regression coefficient βI. Since we do not have any prior information about baseline parameters, α1,λ1,α2, and λ2 prior distributions are assumed to be flat. A widely used prior for frailty parameter σ2 is the gamma distribution with mean one and large variance, G (φφ) say with a small choice of and for the regression coefficients I=1,2,..,k, we use the normal prior with mean zero and large variance ε2. Similar types of prior distributions are used in Ibrahim et al. [14], Sahu et al. [25] and Santos and Achcar [15]. We set hyper-parameters φ=0.0001, ε2=1000. We consider the non informative prior distribution for baseline parameters as the Gamma (1,0.0001).
For both the chains the results were somewhat similar so we present here the analysis for only one chain (i.e. chain 1) for the resulting model. Also due to lack of space we are not providing graphs. Simulated values of the parameters have the autocorrelation of lag k, so every kth iteration is selected as a sample from posterior distribution. The posterior mean and standard error with credible intervals for different sample sizes are reported in Tables 1 and 2 for Model-I, Model-II respectively. From these Tables, it can be observed that the estimated values of the parameters are close to the true values of the parameters and the standard errors decrease as the sample size increases.
To study the Bayesian estimation procedure we use kidney infection data of McGilchrist and Aisbett [4]. The data is regarding recurrence times to infection at point of insertion of the catheter for 38 kidney patients using portable dialysis equipment. For each patient, first and second recurrence times (in days) of infection from the time of insertion of the catheter until it has to be removed owing to infection is recorded. The catheter may have to be removed for reasons other than kidney infection and this regard as censoring. So survival time for a patient given may be first or second infection time or censoring time. After the occurrence or censoring of the first infection sufficient (ten weeks interval) time was allowed for the infection to be cured before the second time the catheter was inserted. So the first and second recurrence times are taken to be independent apart from the common frailty component. The data consists of three risk variables age, sex and disease type GN, AN and PKD where GN, AN and PKD are short forms of glomerulo neptiritis, acute neptiritis and polycyatic kidney disease.
Let T1 and T2 be represents first and second recurrence time to infection. Five covariates age, sex and presence or absence of disease type GN, AN and PKD are denoted by X1, X2, X3, X4, and X5. To analyze kidney infection data success is defined as getting infection first time so we define r=1.
First we check goodness of fit of the T1 and T2. If marginal distributions of T1 and T2 for two proposed distributions fit well then the bivariate distribution of T1 and T2 may be fit well for the same. We used Kolmogorov-Smirnov Goodness-of-Fit Test. Thus from p-values of K-S test we can say that there is no statistical evidence to reject the hypothesis that data are from proposed models in the univariate case and we assume that the models also fit for the bivariate case. Table 3 gives the p-values of Kolmogorov- Smirnov test for the proposed models. Figure 1 shows the parametric versus non-parametric plots.
To analyze kidney data set, various models have been applied by different researchers.
Some of them are, McGilchrist and Aisbett [4] McGilchrist [26], Sahu et al. [25], Boneg [27], Yu [28] and Santos and Achcar [15]. McGilchrist and Aisbett considered semi-parametric Cox proportional hazards model with log-normal frailty distribution and applied Newton-Raphson iterative procedure to estimate the parameters of the model. McGilchrist [26] and Yu [28] both considered the same model as in McGilchrist and Aisbett but McGilchrist estimated the parameters of the model using BLUP, ML and REML methods and Yu proposed modified EM algorithm and penalized partial likelihood method. Sahu et al. considered four parametric models first two are piecewise exponential model with constant baseline hazard say λK within each interval having gamma prior for λK for Model-I and for Model-II normal prior to. Both the models have frailty distribution as gamma. Other two models are multiplicative gamma frailty as Model-III and additive frailty as Model-IV with Weibull as baseline distribution. Santos and Achcar) used MCMC method to estimate the parameters of parametric regression model with Weibull and generalized gamma distribution as baseline and gamma and log-normal as frailty distributions. Boneg considered Cox proportional hazards model and also parametric frailty models. In parametric frailty models he considered Weibull distribution as the baseline and log-normal, Weibull as frailty distributions. He applied MHL and RMHL methods to estimate the parameters of the models.
We run two parallel chains for both models using two sets of prior distributions with the different starting points using Metropolis-Hastings algorithm and Gibbs sampler based on normal transition kernels. On the similar line of simulation, here also we assume same set of prior distributions. We iterate both the chains for 95000 times. We present here the analysis for only one chain with G (a1; a2) as prior for baseline parameters, for both the proposed models. Due to lack of space we are presenting trace plots, coupling from the past plots and sample autocorrelation plots for the parameters of Model I only as shown in Figures 2-4. Gelman-Rubin convergence statistic values are nearly equal to one and Geweke test statistic values are quite small and corresponding p-values are large enough to say the chains attain stationary distribution. Simulated values of parameters have autocorrelation of lag k, so every kth iteration is selected as sample. The posterior mean and standard error with 95% credible intervals for baseline parameters, frailty parameter and regression coefficients are presented in Table 4. The AIC, BIC and DIC values for both the models are given in Table 5. The Bayes factor for the proposed models is also computed.
From the Table 4, we can observe that for these two models, the value zero is not a credible value for the credible interval of the regression coefficient X2 so X2 that is sex variable seems significant. Negative value of β2 indicates that the female patients have a slightly lower risk of infection. The remaining all the covariates X4 i.e. age, X5 i.e. disease type PKD X3 i.e. disease type GN and covariate X4 i.e. disease type AN are insignificant for both the Models.
The estimate of σ2 from two models shows that there is a strong evidence of high degree of heterogeneity in the population of patients. Some patients are expected to be very prone to infection compared to others with same covariate value. This is not surprising, as seen in the data set there is a male patient with infection time 8 and 16, and there is also male patient with infection time 152 and 562.
The comparison between two proposed models is done using AIC, BIC and DIC values given in Table 5. It is observed that both Model I and Model II have AIC, BIC except DIC values are near about same. An alternate way to take the decision about the best model between the proposed Model I and Model II, we use the Bayes factor which is defined as,
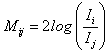
Where Ii is as defined as in equation (7.5)
Using Equation (9.1) we computed Bayes factor M21 which is 6.3162 for our proposed models.
We can observe that the Bayes factor of Model II vs Model I is 6.3162 implies that, there is positive evidence against Model-I, so Model-II is better than Model-I. Thus, Model II is best model of the proposed compound negative binomial frailty models. Now, we are in a position to say that, we have suggested a new shared compound negative binomial frailty model with linear failure rate distribution as the baseline distribution is best for modeling of kidney infection data. For simulation study and to analysis the kidney infection data we used R software.
In the present study we discuss results for the two proposed models of compound negative binomial frailty namely Pareto, linear failure rate distribution as the baseline distributions. Our aim is to find the model which fit best between proposed models. For maximum likelihood estimate, likelihood equations do not converge and method of maximum likelihood fails to estimate the parameters so we use Bayesian approach. Using the Bayesian approach we perform simulation study and analyze kidney infection data. The estimate of frailty variance from different models (Model-I=1.0009; Model-II=1.0001) shows that there is a strong evidence of high degree of heterogeneity in the population of patients. The covariate sex is significant for both models. Negative value of the regression coefficient (β2) of covariate sex indicates that the female patients have a slightly lower the risk of infection. Negative value of the regression coefficient (β5) of covariate, the disease type PKD indicates that the patient with the absence of this diseases have a slightly lower the risk of infection. On the basis of AIC, BIC, DIC and Bayes factor, the Model II, the shared compound negative binomial frailty with linear failure rate distribution is best model for kidney infection.